Journal of Vaccines & Immunization
An International Peer-Reviewed Open Access Journal
ISSN 2053-1273

- Download PDF
- |
- Download Citation
- |
- Email a Colleague
- |
- Share:
-
- Tweet
-

Journal of Vaccines & Immunization
Volume 1, Issue 2, August 2013, Pages 13-21
Original researchOpen Access
Comparison of the live attenuated mumps vaccine (Miyahara strain) with its preattenuated parental strain
- 1 Department of Virology III, National Institute of Infectious Diseases, Musashi-Murayama, 208-0011, Japan
- 2 Department of Pathology, National Institute of Infectious Diseases, Musashi-Murayama, 208-0011, Japan
- 3 Department of Infection Biology, Division of Biomedical Science, Faculty of Medicine, University of Tsukuba, Tsukuba, 305-0006, Japan
- 4 The Chemo-Sero-Therapeutic Research Institute (KAKETSUKEN), Kumamoto, 860-0083, Japan
*Corresponding author: Atsushi Kato, Division of Radiological Protection and Biology, National Institute of Infectious Diseases, 1-23-1 Toyama, Shinjuku-ku, Tokyo, 162-8640, Japan, Tel.: 81 3 5285 1111, ext. 2421; Fax: 81 3 5285 1194. E-mail: akato@niid.go.jp
Received 3 February 2013 Revised 9 May 13 Accepted 17 May 2013 Published 24 May 2013
DOI: http://dx.doi.org/10.14312/2053-1273.2013-3
Copyright: © 2013 Nagata S, et al. Published by NobleResearch Publishers. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited
AbstractTop
Live attenuated mumps vaccines have been developed by passaging the field isolates in cells that differ from the natural host. To study mumps viral (MuV) attenuation at the genomic level, we compared a live attenuated mumps vaccine (Miyahara strain) to its preattenuated parental strain. These two strains exhibited several phenotypic differences. The parental strain grew faster, reached a higher titer, and formed larger plaques and syncytia in Vero cells compared to the vaccine strain. In addition, intracranial injection of parental strain in neonatal rats resulted in greater ventricular enlargement than that caused by the vaccine strain. Four nucleotide changes leading to amino acid substitutions were found between the two viral genomes. One change is present in the N and L genes, respectively, and two in the F gene. The fusogenic activity of the cloned parental F gene evaluated by a cell-cell fusion assay was weaker than that of the vaccine F gene, and did not correspond to the activity caused by the living parental and vaccine strains. The transcriptional activities of N and L proteins were monitored by a CAT minigenome assay. Cloned parental N gene produced almost the same CAT activity as vaccine N, and cloned parental L gene produced significantly higher CAT activity than vaccine L. Thus among four nucleotide changes, the three occurring in N and F were not found to relate to the viral outcome, but we confirmed that at least one change in the L protein was involved in the growth phenotype of parental MuV.
Keywords: attenuation; fusion; mumps virus; neurovirulence; replication; vaccine
IntroductionTop
Mumps is an acute, contagious vaccine-preventable disease caused by the mumps virus (MuV). A mumps epidemic occurs every 3 to 5 years in unvaccinated populations, and humans are the only natural host for MuV infection. MuV is transmitted by the respiratory route and replicates primarily in the nasal epithelium, leading to viremia, and it triggers secondary viral replication [1]. Up to 30% of MuV infections are asymptomatic or show only nonspecific respiratory symptoms. In 70% to 90% of symptomatic infections, MuV invasion of glandular epithelium results in mumps parotitis. MuV enters the cerebrospinal fluid (CSF) during the viremia. CSF pleocytosis leads to a form of aseptic meningitis that occurs in about 10% of mumps cases. More serious complications, such as deafness and encephalitis, occur at a low frequency [1].
MuV belongs to the genus Rubulavirus of the family Paramyxoviridae and has a negative-strand nonsegmented RNA genome of 15,384 bases with seven genes encoding eight proteins: the nucleocapsid (N), V, phospho (P), matrix (M), fusion (F), small hydrophobic (SH), hemagglutinin-neuraminidase (HN), and large (L) proteins. The N, P, and L proteins participate in viral replication and, together with the genomic RNA, form the ribonucleocapsid (RNP). In the virion, RNP is surrounded by a host-derived membrane undercoated by the M protein. The HN and F proteins projected onto the virion surface are responsible for cell binding and the entry of MuV. The functions of V and SH are less well understood [2–5]. The SH gene sequence is variable and is therefore used as the basis of MuV genotyping [6–8].
Live attenuated MuV vaccines have been developed by adapting the virus for different species of host or for different conditions of temperature, after repeated trial and error. The first vaccine, Jeryl Lynn strain, was introduced into the USA in 1967 and contributed to a dramatic reduction in mumps infection rates. To date, more than 10 vaccines have been produced. Some vaccines were found to retain a limited pathogenicity [9–11], and some tend to lose immunogenicity because of over-attenuation [12]. Five vaccines were developed in Japan (Urabe [13], Hoshino [14], Torii, Miyahara and NK-M46), and they were found to cause aseptic meningitis, although much less frequently than the natural infection rate [15, 16]. The virulence of MuV has recently been studied using the reverse genetic techniques established for Jeryl Lynn strain cDNA [17]. The genetic basis for attenuation has been vigorously studied [18–23], but a common genetic factor for MuV attenuation has not yet been identified.
In this study, we compared a Japanese mumps vaccine, Miyahara strain, with its parental strain in the preattenuated position to investigate the molecular basis of attenuation. The parental strain apparently has a different phenotype. The full genome sequences of the parental strain and the vaccine revealed four nucleotide substitutions. Those changes accompany the amino acid substitution, and one amino acid change occurring in the L protein was linked to replication capacity in vitro.
Materials and methodsTop
Viruses
The MuV Miyahara isolate was originally obtained from a mumps patient in 1970 in Japan. A passage history of the Miyahara strain is indicated in Figure 1. The Miyahara field isolate had been lost, but the strain that locates on the preadapted position of chicken embryo fibroblasts (CE) passages (exact position was not known) was retrieved from the freezer of the National Institute of Infectious Diseases, Japan and used in this study as a parental strain with the approval of the Kaketsuken (the Chemo-Sero-Therapeutic Research Institute, Kumamoto, Japan). We purchased the live attenuated Miyahara MuV vaccine (lot 305) from the Chemo-Sero-Therapeutic Research Institute. Wild-type MuV Odate-1 [24, 25] and 02-49 were kindly provided by Dr. H. Saito (Akita Prefectural Institute of Public Health, Akita, Japan) and Dr. Y. Matsui (Niigata Prefectural Institute of Public Health and Environmental Science, Niigata, Japan), respectively. In all virus stocks used in this study, defective interfering particles that might influence the results were not found.
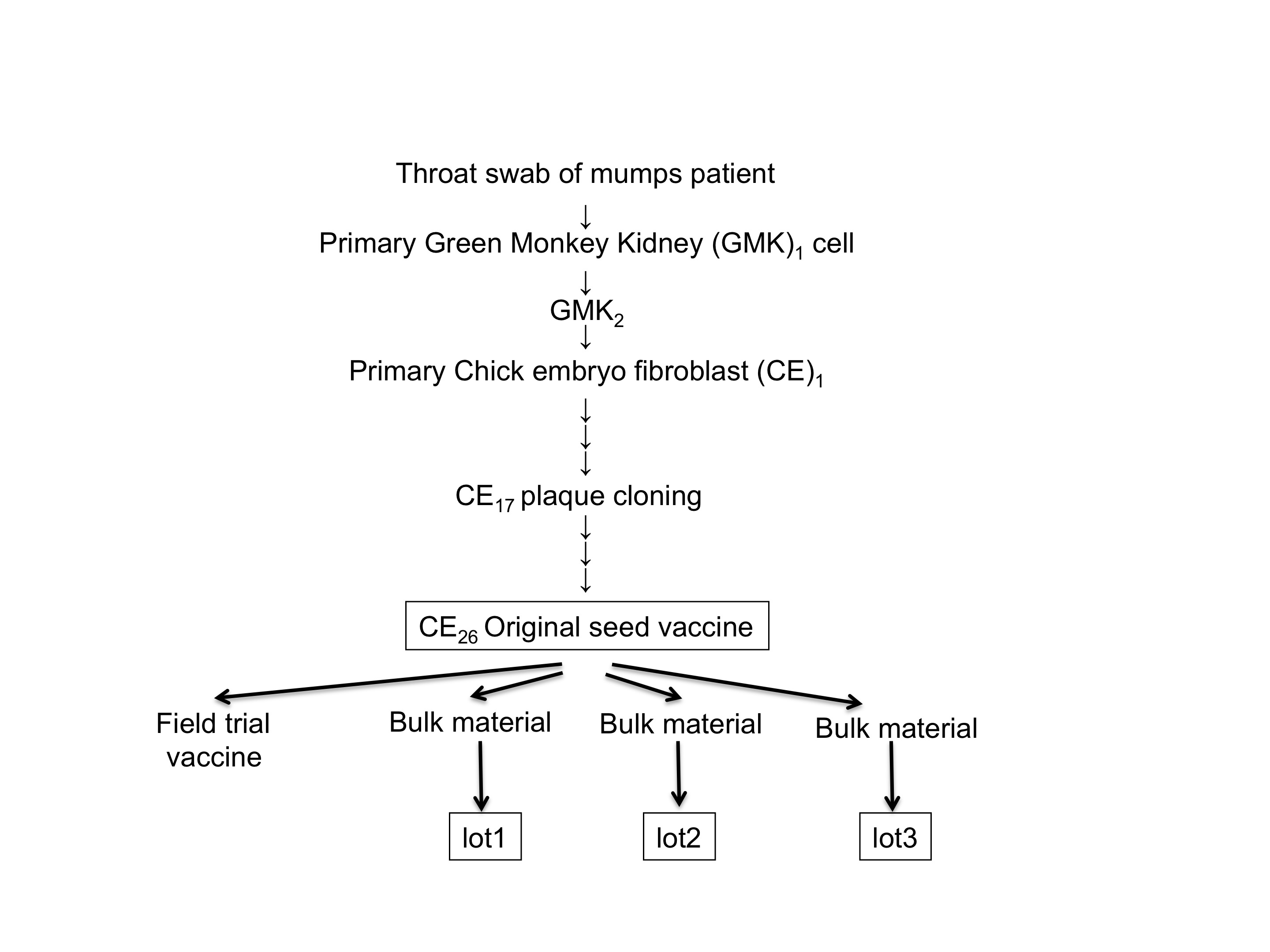
Figure 1 Passage history of the MuV Miyahara strain. MuV originally isolated from a throat swab of a 4-year-old female mumps patient in primary green monkey kidney cells is shown as GMK1. Another passage in the same GMK cells is shown as GMK2. Passages in the primary cell culture of chick embryo fibroblasts are shown as CE. At the 17th passage of CE (CE17), MuV was plaque purified. The test lot produced from CE26 was administered as the field trial vaccine. CE26 was licensed as the original seed for a live attenuated mumps vaccine, the Miyahara strain, in 1985 in Japan. The exact position of the Miyahara parental strain used in this study is not mapped due to the loss of records. Three bulk materials (lots 1, 2, and 3) have been produced, and the vaccine (lot 305) used in this study was produced from bulk lot 3.
Cell cultures and virus infection
Vero cells were grown in minimal essential medium (MEM) supplemented with 5% (v/v) bovine serum (BS) and inoculated with parental or vaccine strains at a multiplicity of infection (m.o.i.) of 0.01 cell infection unit (CIU)/cell unless otherwise noted. BHK-T7 cells [26] were kindly supplied by Dr. N. Ito (Gifu University, Gifu, Japan), and were maintained in Glasgow-MEM (Life Technologies Japan, Tokyo) containing 5% fetal BS.
For the growth kinetics study of parental and vaccine strains, 0.1 mL of culture supernatant was harvested daily until 5 days post-infection. The virus titer was then measured at least five times in each sample. MuV strains were titrated using Vero cells. Briefly, a diluted series of MuV was inoculated onto confluent cells in a 12-well plate after the removal of medium. After adsorption for 1 h, 1 mL of medium was added to each well. Thirty-six hours later, infected cells were treated with rabbit anti-MuV hyperimmune serum followed by fluorescein isothiocyanate (FITC)-conjugated anti-rabbit immunoglobulin G. FITC-positive cells were measured, and the virus titer (CIU/ml) was calibrated.
For plaque formation, MuV was inoculated onto confluent Vero cells in a 6-well plate. After adsorption for 1 h, 1.0% (w/v) agarose in MEM containing 0.5% BS was overlaid on the cell culture and incubated at 37°C for 8 days. An agarose in MEM containing 0.01% (w/v) Neutral Red solution (Sigma-Aldrich Japan, Tokyo) was again overlaid onto the first agarose layer and incubated for several hours until cells were stained. For the syncytium formation, MuV was inoculated onto cells at an input m.o.i. of 0.01 CIU/cell. Two days later, syncytia were observed by treating the cells with rabbit anti-MuV hyperimmune serum followed by FITC-conjugated anti-rabbit immunoglobulin G. FITC-positive fused cells were then counted.
Neurovirulence assay
Fertilized Lewis rats were purchased from Charles River Laboratories Japan (Yokohama, Japan). One-day-old littermates were inoculated intracranially with 20 µL of each MuV strain containing 100 CIU or phosphate-buffered saline (PBS) and raised with their respective dams. This experiment was performed according to the Institutional Animal Care and Use Committee regulations (Permission number: 211072). Under isoflurane anesthetic inhalation, these rats were sacrificed 30 days after the inoculation. Their heads were removed and fixed in 10% buffered formalin solution. Hydrocephalus, characterized by the excessive accumulation of CSF in the cerebral ventricles, was scored as described [27]. The sizes of the brain and lateral ventricles in a digital picture were measured using ImageJ software (National Institutes of Health, Bethesda, MD). The neurovirulence score for each strain is indicated by the mean score for all brains within an inoculated group. We analyzed differences in the scores using Student’s t-test as described [19, 27].
PCR cloning and sequence analysis
The genome of MuV was extracted from the viral fluid using the High Pure Viral RNA Kit (Roche Diagnostics, Tokyo). Extracted RNAs were reverse-transcribed and amplified using the PrimeScript One Step RT-PCR Kit (Takara Bio, Otsu, Japan) with five sets of specific primer pairs that cover the entire sequence of the Miyahara vaccine strain as registered in the DNA database (accession number AB040874). These primer pairs amplified five regions that partially overlapped each other: 1(1-4592, the numericals in parentheses indicate the nucleotides from the 3’ end), 2(3051-7413), 3(6049-10233), 4(8516-12935), and 5(11328-15384).
The amplified fragments were cloned into the pCR2.1 vector (Life Technologies Japan), yielding pCR-1, pCR-2, pCR-3, pCR-4, and pCR-5, respectively. We sequenced plasmids obtained from three independent colonies of transformed E. coli. Both termini of the genomes were amplified using the 5’ and 3’ RACE System (Life Technologies Japan). Nucleotide sequences were read using a BigDye Terminator v3.1 cycle sequencing kit with a 3130xl Genetic Analyzer (Life Technologies Japan). The final sequence was determined if the three sequences obtained from independent clones were identical. If these were not concordant, sequences were determined by the RT-PCR sequencing of bulk RNA. Obtained sequences were registered in the DNA database (accession number AB744048 and AB744049 for Miyahara vaccine lot 305 and parental strain, respectively).
The sequence of the Miyahara vaccine (AB744048) was not identical to the previously registered sequence (AB040874); it had a few nucleotide changes. We ignored these changes in our further studies as an error of the previous sequence, because these nucleotides were not found in the vaccine (AB744048) or in the parental strain (AB744049). The N, P, and L genes of each strain were cloned into the downstream site of the T7 promoter in a pGEM-3 vector (Promega, Tokyo), yielding pGEM-N, pGEM-P, and pGEM-L. The F and HN genes were cloned into the downstream site of the EF-BOS promoter in the pKS336 vector (AF403737, kindly provided by Dr. K. Sakai, National Institute of Infectious Diseases, NIID, Japan), yielding pKS-F and pKS-HN.
Fusion assay
One microgram each of pKS-F, pKS-HN, and pDsRed plasmids (Clontech, Takara Bio, Shiga, Japan) was cotransfected into confluent Vero cells using FuGENE 6 (Promega, Tokyo). Transfected cells were cultured for 24 h at 37C or 39.5C. Fused cells were observed by inverted fluorescent microscopy (AxioVert, Carl Zeiss Japan, Tokyo) and digitally captured using a CCD camera (AxioCam, Carl Zeiss Japan). The sizes of fused cells in the digital picture were scored using ImageJ software.
CAT minigenome assay
BHK-T7 cells, a baby hamster kidney (BHK) cell line stably expressing T7 polymerase [26], were seeded in 12-well plates at the semi-confluent level. Unless otherwise noted, 0.2 µg of pGEM-N, 0.2 µg of pGEM-P, 0.8 µg of pGEM-L, and 1.0 µg of pMuV-CAT were transfected using Lipofectamine 2000 (Life Technologies Japan). The plasmid pMuV-CAT is designed to produce a MuV-negative sense minigenome that codes the chloramphenicol acetyltransferase (CAT) reporter gene at an antisense manner [17]. CAT activity was assayed using the FAST CAT Chloramphenicol Acetyltransferase Assay Kit (Molecular Probes, Eugene, OR). Samples and the CAT reference standard were applied on a silica thin layer chromatography plate (Whatman, GE Healthcare Japan, Tokyo) and developed with chloroform:methanol (9:1) solvent.
Acetylation images were captured digitally using the luminescent image analyzer LAS-3000 (Fujifilm, GE Healthcare Japan). CAT activities were semiquantitatively indicated as a score by adding up the digital intensity of acetylated products at hydroxyl position 1 or 3 of chloramphenicol (CAP) and twice the intensity of those acetylated at both 1 and 3 hydroxyl positions, using ImageJ software. The experiments were performed in triplicate, basically as described [17] except for our use of the non-radioisotope detection method.
Results and discussionTop
Characteristics of the Miyahara parental strain and vaccine in cells
Live attenuated MuV vaccines have been developed by adapting the virus for different species of host or for different conditions of temperature, after repeated trial and error. Five vaccines were developed in Japan through the adaptation to CE. During the vaccines’ creation, candidates were designed to have the characteristic marker producing a smaller plaque than the parental field isolate in Vero cells. We compared the phenotypes of the preattenuated Miyahara parental strain to the live attenuated Miyahara commercial vaccine (Figure 1). The plaques caused by the parental strain were larger than those produced by the vaccine (Figure 2A). The parental plaque consisted of a clear edge and a colorless inside, but the vaccine plaque was unclear and turbid. This morphological character corresponded well with the developmental marker of the vaccine.
Differences in plaque sizes often reflect the viral capacity for fusogenicity, growth, or cytopathogenicity. We thus compared the syncytia of the two viruses at two days post-infection by staining the infected Vero cells with the anti-MuV antibody. The parent strain showed larger syncytia in Vero cells than the vaccine did (Figure 2B). Viral growth at the multistep condition (m.o.i. of 0.01) was then compared at three different temperatures (32°, 37°, and 39.5°C). The results from a representative of three independent experiments runs are shown in Figure 2C. At 32°C, the parental strain and the vaccine grew at almost the same growth rate.
In the higher temperature conditions, however, the parental strain increased faster than the vaccine and reached a higher titer compared to the vaccine. The virus titers of the parental strain at 3 days post-infection were 8.0×105 (32°C), 7.9×106 (37°C), and 3.0×107 (39.5°C) CIU/mL (Figure 2C, filled black arrows). These results indicated the increased growth phenotype of the parental strain with increasing temperature. In contrast, the vaccine grew almost identically in the three temperature conditions and reached identical titers (3.8×106 at 32°C, 2.7×106 at 37°C, and 4.3×106 CIU/mL at 39.5°C) at 5 days post-infection (Figure 2C, outlined white arrows). We tried to compare the viral growth at 42°C but failed to obtain growing viruses from the culture supernatant because of the high-temperature sensitivity of the Vero cells (data not shown).
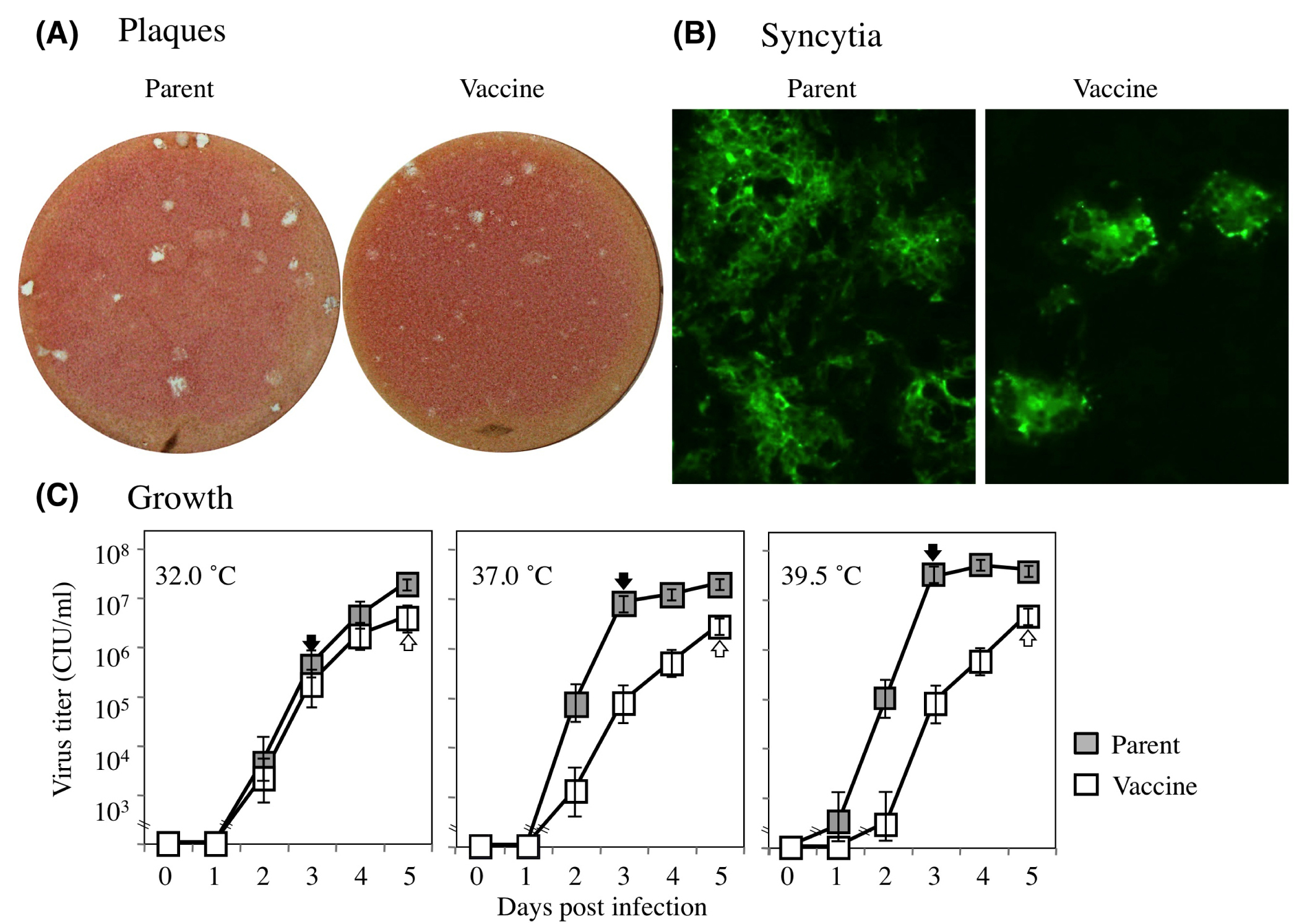
Figure 2 Phenotypic comparisons of parental strain and vaccine of MuV. Plaques produced on Vero cell culture (A). Syncytia found in Vero cell culture (B). Syncytia were reacted with rabbit anti-MuV hyperimmune serum as described in the materials and methods section. The growth of parental MuV and vaccine was compared under the temperature conditions indicated (C). A representative of three independent studies (32° and 37°C, 37° and 39.5°C, and 32° and 39.5°C) is shown. Closed boxes show the titer of parent MuV; open boxes show that of vaccine MuV. Each point indicates the mean CIU/mL titer of six calculations. Error bars: SEM..
Characteristics of neurovirulence in rats
Since humans are the only natural host for MuV, no routine method for assaying the pathogenicity of MuV has been established, although attempts have been made with the Cynomolgus monkey [28, 29], marmoset [30], and hamster [31]. We adopted a neurovirulence assay model using neonatal rats [27, 32]. In the present study, we injected two field isolates (Odate-I and 02-49) and PBS intracranially into littermates of rats as the positive and the negative controls along with the parental strain and vaccine (Figure 3). Odate-I was isolated from a meningitis patient during a mumps outbreak in 1993 with a high incidence of aseptic meningitis [24, 25], while 02-49 was isolated from a parotitis patient in 2002 in Japan.
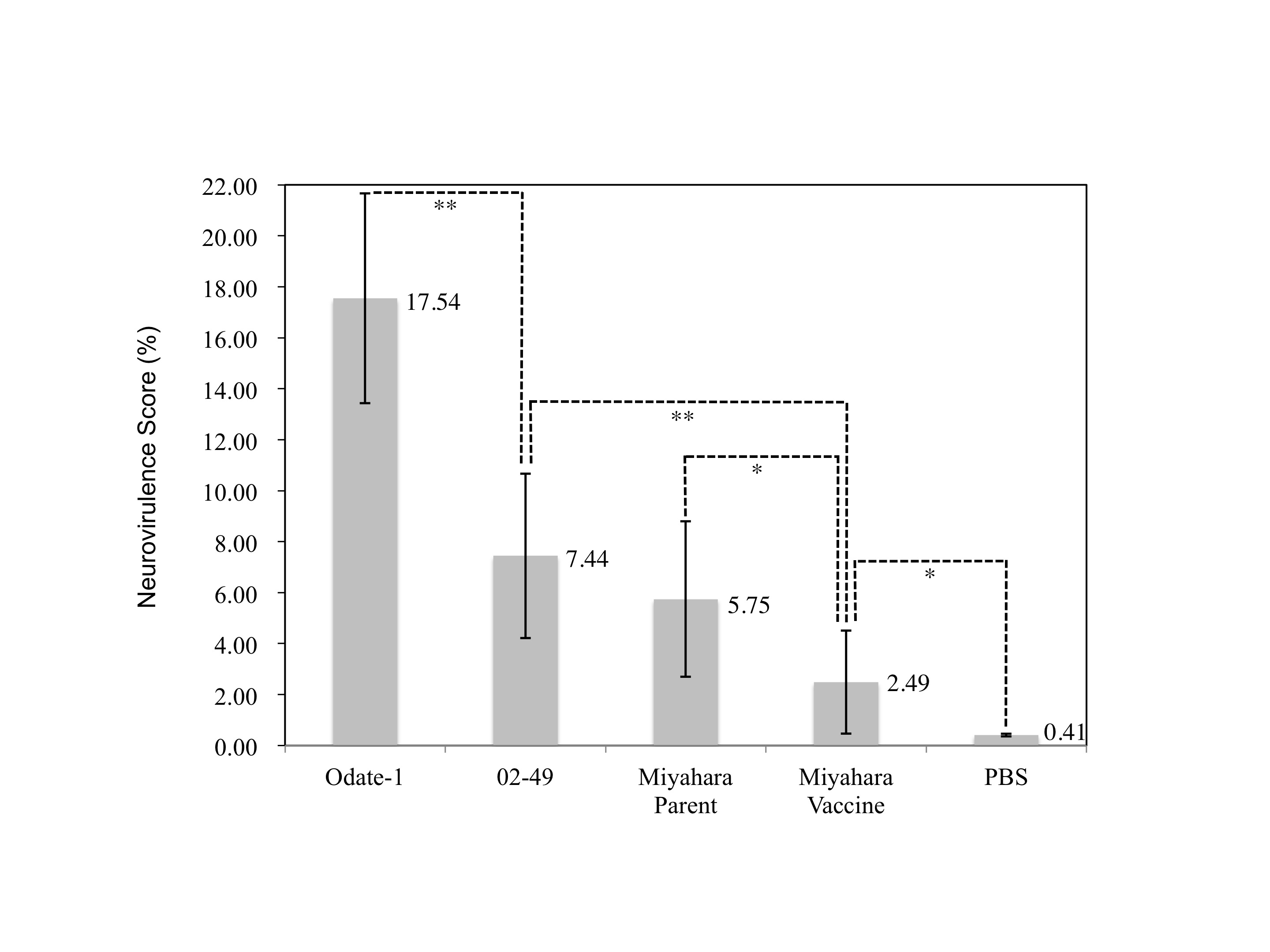
Figure 3 The severity of hydrocephalus in rats inoculated with the parental strain and vaccine of MuV. We tested the neurovirulence of parental MuV and vaccine as described in the materials and methods section. All bars and numerals represent the mean values of neurovirulence scores. Error bars: SEM. **, P<0.01; *, P<0.05). The Miyahara vaccine’s score was 2.49±2.41(n=20), significantly lower than that of 02-49 (P=0.006<0.01) but higher than that of PBS (P=0.026<0.05).
The neurovirulence score for Odate-I was 17.54±4.12 (n=18) and for 02-49 was 7.44±3.22 (n=18). The decrease of neurovirulence observed in 02-49 was statistically significant (P<0.01). PBS-inoculated rats did not show any apparent changes and the score was 0.41±0.05 (n=24). The Miyahara vaccine’s score was 2.64±2.41 (n=20), significantly lower than that of 02-49 (P<0.01), indicating the avirulent phenotype of approved vaccine. The neurovirulence score of the parental strain was 5.75±3.05 (n=18). This score was higher than that of the vaccine (P=0.048<0.05), but no significant difference between the parental strain score and that of 02-49 was revealed (P=0.436). This result is in line with the finding that the Miyahara parental strain was moving toward adapting to CE cells and thus retaining the neurovirulence.
Differences in genome sequences
At this point, we had found some biologically significant differences between the Miyahara parent strain and the vaccine. A comparison of the genome sequence between the two strains was then necessary to better understand their biological functions at the molecular level. We cloned three independent fragments amplified from each genome and sequenced them carefully. Both termini were confirmed using the 3’ and 5’ RACE methods as described in the materials and methods section. Our analysis of the sequence data (AB744048 and AB744049) yielded four nucleotides that differed between the two strains: 1337 in the N gene, 5024 and 5129 in the F gene, and 14355 in the L gene (the numerals indicate the position from the 3’ end). There was no deletion or insertion in the genome, and no nucleotide exchange had occurred in the noncoding region. All four exchanges were accompanied by the amino acid changes 1337N398(Val → Ile), 5024F160(Gln → Arg), 5129F195(Phe → Ser), and 14355L1973(Phe → Ser).
Fusogenicity of cells transfected with cloned F and HN genes
The F protein of MuV is responsible for the virus-cell and the cell-cell membrane fusion with the cooperation of the HN protein. To determine whether the amino acid exchanges in the F protein facilitated the fusion activity, we cotransfected plasmids coding each F and HN protein into Vero cells together with the pDsRed, and incubated them at 37°C or 39.5°C. The syncytia thus formed emitted red fluorescence (Figure 4). In marked contrast to the syncytia with live MuV-infected cells (Figure 2), the sizes of the syncytia associated with the parental F gene were smaller than those associated with the vaccine F gene.
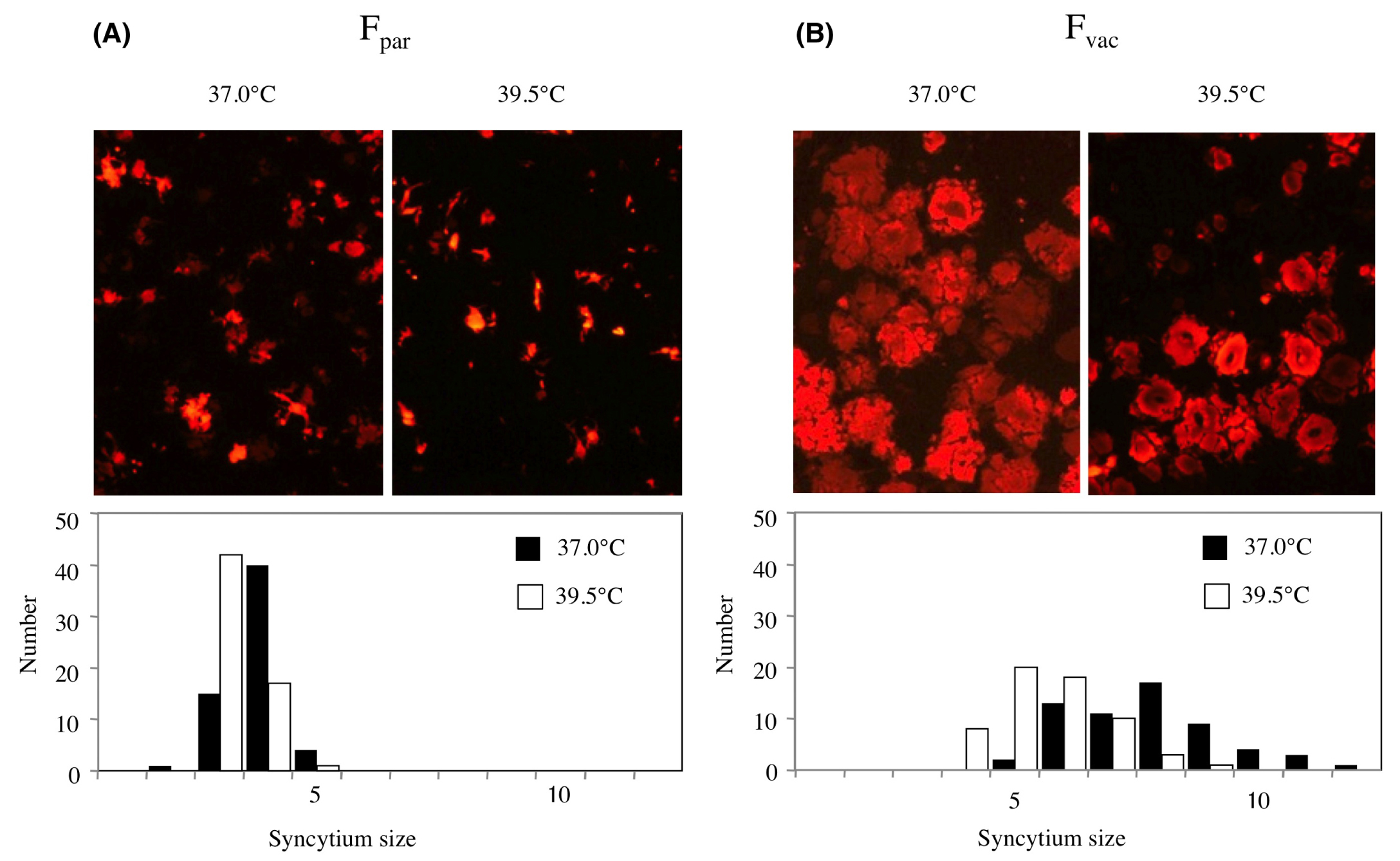
Figure 4 Syncytium formation assay by cloned F and HN genes. Vero cell cultures were transfected with plasmids encoding cloned F gene from parent (Fpar) or vaccine (Fvac) and HN gene together with pDsRed. Fused cells emitting fluorescence are shown. At 37°C (closed bar) or 39.5°C (open bar), the images of syncytia were digitally taken and the diameters were plotted as histograms.
The diameters of 50 randomly taken syncytia were measured and the histogram was drawn at 37°C and 39.5°C. The syncytia produced at 39.5°C were smaller than those obtained at 37°C. The temperature-independent syncytia formation by cloned F genes was the opposite of the higher growth rate of living parent MuV at 39.5°C compared to 37°C. These results showed that the vaccine phenotype was not explained by 5024F160(Gln → Arg) and 5129F195(Phe → Ser) mutations.
Minigenome activity of cloned genes
The P and L proteins form the viral RNA polymerase and the N protein surrounds the genomic RNA to form an active template. To evaluate the relevance of 1337N398 (Val → Ile) and 14355L1973(Phe → Ser) in vitro, we used a minigenome in which all MuV genes of the Miyahara strains were replaced with a CAT gene. CAT activity reflected the amount of CAT mRNA produced from the minigenome. We found no CAT activity in the experimental conditions when one of three proteins was depleted, but found that the use of 0.2 µg, 0.2 µg, and 0.4 µg of N, P, and L plasmids, respectively, resulted in the optimum CAT activity. Inputting the vaccine N (Nvac) or parental N (Npar) plasmid from 0 to 0.1 and 0.2 µg, the three clearer forms of acetylated products (1-, 3-, and 1,3-chloramphenicol (CAP)) were observed (Figure 5A). However, further input of the N plasmid (0.4 and 0.6 µg) did not result in an increase in acetylated products but rather their decrease. Maximum CAT activities were shown at 0.2 µg of N plasmid, and were almost the same between the two N plasmids. By the digital intensity of acetylated products, the amount of products was indicated as 34.1 (vaccine) and 35.1 (parent). These results showed that Nvac and Npar had no functional difference and that both were equivalent in the CAT assay. Any relevance of 1337N398(Val → Ile) was not identified under this condition.
We then added various amounts of vaccine L (Lvac) or parental L (Lpar) plasmid together with N and P plasmids. Too many or too few L plasmids did not result in high CAT activity (Figure 5B). The maximum CAT activity was found under the conditions in which 0.4 µg or 0.8 µg of Lpar or Lvac was added. The digital intensity of acetylated products was indicated as 51.8 (parent) and 32.8 (vaccine). Higher CAT activity was observed in Lpar-transfected cells than in Lvac-transfected cells. These results indicate that Lpar and Lvac did not have the same effect and that Lpar had 1.6-fold (51.8/32.8) higher transcription and replication capacities. We tested and confirmed this higher capacity of Lpar to Lvac by measuring acetylated products in three independent experiments. Thus, 14355L1973(Phe → Ser) was relevant under this condition.
Although an effect of the N mutation was not observed in the above experiment, it is possible that the mutation in the N gene might have an effect in combination with the mutation in the L gene. We therefore added various amounts of Lvac with a combination of fixed amounts of Nvac, and we also added Lpar with a combination of fixed amounts of Npar. Higher CAT activity was observed in Npar/Lpar-transfected cells than in Nvac/Lvac-transfected cells. Maximum CAT activities were obtained when 0.4 µg or 0.8 µg of Lpar or Lvac was added. The scores shown by the digital intensity of acetylated products were 57.4 (parent) and 35.4 (vaccine) (data not shown). These results again showed the importance of 14355L1973(Phe → Ser), but in a comparison of Nvac/Lvac and Npar/Lpar (57.4/35.4 = 1.6 fold) to Nvac/Lvac and Nvac/Lpar (51.8/32.8 = 1.6 fold, Figure 5B), the fold ratio of vaccine to parent gave the same value. Similar results were also obtained when various amounts of Nvac were used in combination with fixed amounts of Lvac and when Npar was used in combination with fixed amounts of Lpar (data not shown). These findings show no contribution of 1337N398 (Val → Ile) mutation on CAT activity in combination
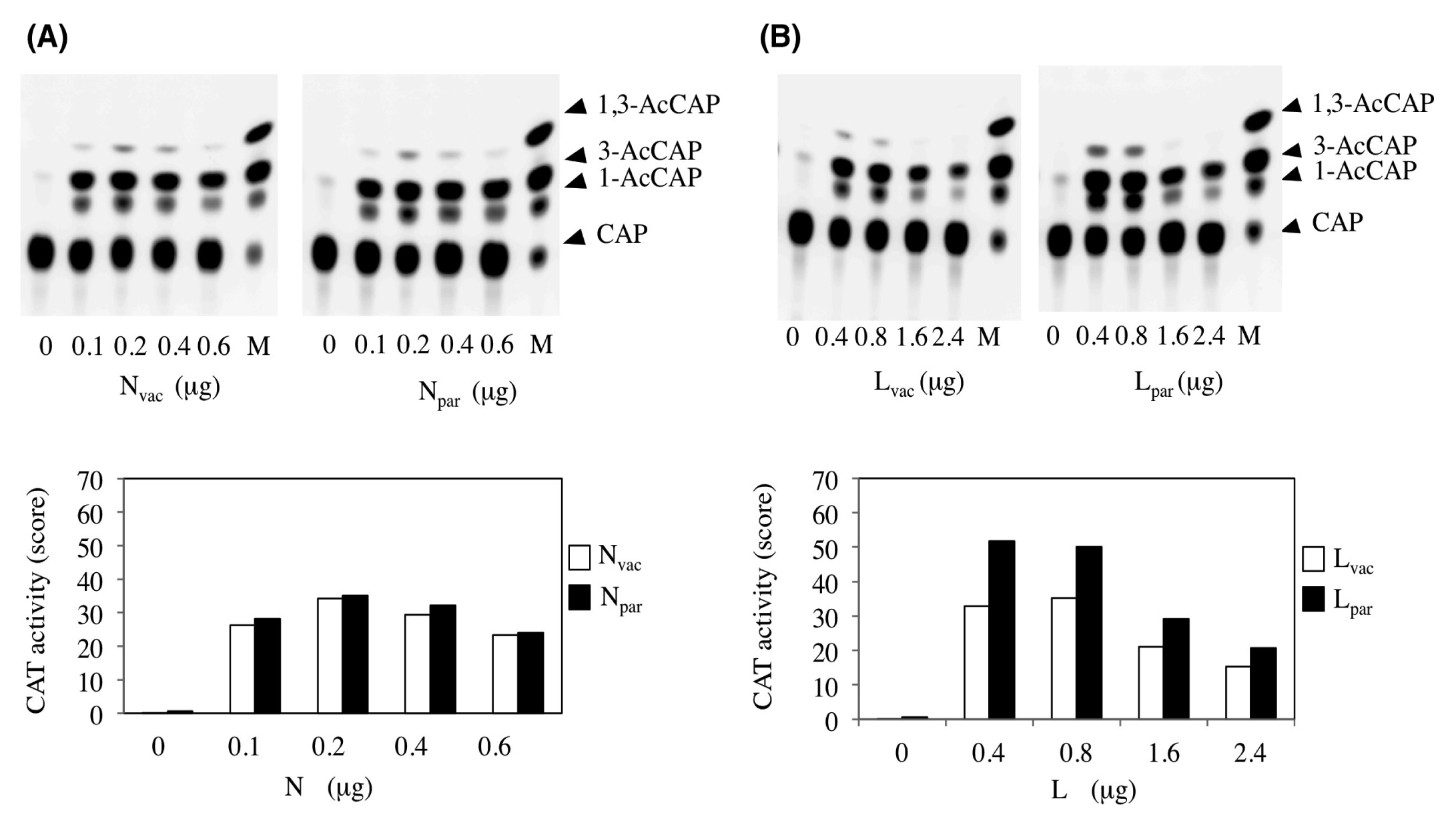
Figure 5 CAT minigenome assay for evaluating mutations in the N and L genes. Various amounts of the N gene from parent (Npar) or vaccine (Nvac) were transfected into the cells together with P and vaccine L (Lvac) genes and the CAT minigenome, as described in the materials and methods section (A). Various amounts of the L gene from parent (Lpar) or vaccine (Lvac) were transfected together with P and Nvac genes and the CAT minigenome (B). The acetylated products developed in TLC plates are shown in the upper panel. The intensities of acetylated spots obtained from the digital photograph are indicated as the CAT activity bar graphs in the bottom panel. Open and closed bars show the results obtained by vaccine- and parent-derived gene transfection, respectively. Data are from a representative experiment performed in triplicate.
DiscussionTop
Wild-type MuV exhibits neurovirulence, and even approved attenuated vaccines carry some risk of neurovirulence and can cause aseptic meningitis, although the frequency of its occurrence with the mumps vaccine is low compared to the natural infection. Approximate frequencies are specifically determined by data from administered vaccine strains [1, 9], which indicate some genetic factor dictating MuV’s neurovirulence. The identification of such a gene or genes is thus important for developing a safe vaccine.
It was reported that the Urabe mumps vaccine is a mixture of viruses differing at amino acid 335 of the HN gene, and one form was associated with neurovirulence [33]. Some virological studies supported this finding [21, 22]. However, the complete nucleotide sequences of several stocks of the Urabe strain did not support this finding [34, 35], and reverse genetic work finally showed that HN335 was not a crucial determinant of Urabe neurovirulence [36].
The Jeryl Lynn vaccine strain adapted to human neuronal cells had three amino acid substitutions in N468(Phe → Pro), M85(Val → Ala) and L1165(Glu → Asp), and the wild-type 88-1961 strain adapted to CE fibroblasts had three amino acid substitutions in F91(Ala/Thr → Thr), HN466(Ser → Asn) and L736(Ile → Val) [37]. Swapping of the respective gene from the Jeryl Lynn strain to the 88-1961 strain resulted in a gradual attenuation with the increase in the number of genes and yielded no critical gene for viral virulence, and swapping those from the wild-type to the Jeryl Lynn strain resulted in a gradual virulence to the contrary. [23]. These results suggest that there is no specific gene determining viral attenuation and neurovirulence, and that viral attenuation and neurovirulence are determined by a sum or a certain combination of noncritical genes in the context of viral life cycle.
We started to study MuV attenuation using one of the Japanese vaccines, the Miyahara strain, and its preattanuated parental strain. We found that the plaque size, viral growth, temperature sensitivity, fusogenicity and neurovirulence of the two strains were different (Figurs 2 and 3). The genome sequences of the parental strain and vaccine were carefully determined by two-step experiments. In the first step, three independent cDNA clones of both strands were sequenced and the sequence was determined only when the results were identical. In a few non-concordant cases, sequences were determined by the RT-PCR sequencing of bulk RNA as a second step. The results indicated that each virus source was highly homogeneous.
Considering that the Jeryl Lynn vaccine contains JL2 and JL5 component stains which differ by over 400 nucleotides [38], it was surprising for us that these phenotypes were probably caused by four nucleotide exchanges between the genomes. Among the four exchanges, 5024F160(Gln → Arg) and 5129F195(Phe → Ser) seemed not to be related to the Miyahara phenotype but were epiphenomena (Figure 4). Moreover, 5129F195(Phe) is found among known mumps strains independent of vaccines or field isolates. For example, the Odate-1 strain used for the rat neurovirulence test as a virulent strain in the present study has 5129F195(Ser), while the Urabe strain approved for the mumps vaccine in Japan has 5129F195(Phe). The relevance of 5129F195(Phe) to low fusogenic activity has been reported in other strains [39, 40]. The discrepancy in fusogenic activity between the living virus and the cloned F gene was not resolved in this study.
ConclusionTop
We found that 14355Lpar1973(Phe) resulted in higher CAT activity than 14355Lvac1973(Ser), but that 1337Npar398 (Val → Ile) mutation did not result in a significant difference (Figure 5). We concluded that an amino acid substitution occurring in 14355Lpar1973(Ser) is associated with attenuation by reducing replication and transcription activity. Six conserved L domains (I-VI) are known between two different families of Mononegavirales, rhabdoviridae and Paramyxoviridae, and are postulated to constitute the specific enzymatic activities [41]. However, the amino acid change observed herein in L1973 does not belong to any of the conserved regions. How the amino acid change in L1973 might alter transcriptional activity is unknown. Another possibility also remains that the mutations contribute to viral attenuation in a way that the fusogenic or the minigenome assay in vitro does not capture. The importance of amino acid mutations during the attenuation process will be revisited as a next step using the reverse genetics technique in the context of viral multiplication in cells and of viral neuropathogenecity for neonatal rats.
Acknowledgements
We thank Y. Ami and Y. Suzaki (NIID) for their helpful support with animal experiments, M. Tahara (NIID) for providing the T7-expressing cells, and T. Kubota (NIID) for the helpful advice.
Funding
This work was supported by a grant from the Ministry of Health, Labour and Welfare, Japan.
Conflict of interest
All the authors declare that they have no conflict of interest.
ReferencesTop
[1] Rubin SA, Carbone KM (2011) Mumps, In: Longo DL, Fauci AS, Kasper DL, Hauser SL, Jamenson JL, Loscalzo J, eds. Harrison's Principles of Internal Medicine. 18th ed. New York: The McGaw Hill Companies Inc., p1608–1610.
[2] Malik T, Shegogue CW, Werner K, Ngo L, Sauder C, et al. (2011) Discrimination of mumps virus small hydrophobic gene deletion effects from gene translation effects on virus virulence. J Virol 85:6082–6085. Article Pubmed
[3] Takeuchi K, Tanabayashi K, Hishiyama M, Yamada A (1996) The mumps virus SH protein is a membrane protein and not essential for virus growth. Virology 225:156–162. Article Pubmed
[4] Xu P, Li Z, Sun D, Lin Y, Wu J, et al. (2011) Rescue of wild-type mumps virus from a strain associated with recent outbreaks helps to define the role of the SH ORF in the pathogenesis of mumps virus. Virology 417:126–136. Article Pubmed
[5] Xu P, Luthra P, Li Z, Fuentes S, D'Andrea JA, et al. (2012) The V protein of mumps virus plays a critical role in pathogenesis. J Virol 86:1768–1776. Article Pubmed
[6] Jin L, Rima B, Brown D, Orvell C, Tecle T, et al. (2005) Proposal for genetic characterisation of wild-type mumps strains: preliminary standardisation of the nomenclature. Arch Virol 150:1903–1909. Article Pubmed
[7] Kidokoro M, Tuul R, Komase K, Nymadawa P (2011) Characterization of mumps viruses circulating in Mongolia: Identification of a novel cluster of genotype H. J Clin Microbiol 49:1917–1925. Article Pubmed
[8] Takeuchi K, Tanabayashi K, Hishiyama M, Yamada A, Sugiura A (1991) Variations of nucleotide sequences and transcription of the SH gene among mumps virus strains. Virology 181:364–366. Article Pubmed
[9] Bonnet MC, Dutta A, Weinberger C, Plotkin SA (2006) Mumps vaccine virus strains and aseptic meningitis. Vaccine 24:7037–7045. Article Pubmed
[10] Rafiefard F, Johansson B, Tecle T, Orvell C (2005) Characterization of mumps virus strains with varying neurovirulence. Scand J Infect Dis 37:330–337. Article Pubmed
[11] Rubin SA, Afzal MA (2011) Neurovirulence safety testing of mumps vaccines--historical perspective and current status. Vaccine 29:2850–2855. Article Pubmed
[12] Ong G, Goh KT, Ma S, Chew SK (2005) Comparative efficacy of Rubini, Jeryl-Lynn and Urabe mumps vaccine in an Asian population. J Infect. 51:294–298. Article Pubmed
[13] Yamanishi K, Takahashi M, Kurimura T, Ueda S, Minekawa Y, et al. (1970) Studies on live mumps virus vaccine. 3. Evaluation of newly developed live mumps virus vaccine. Biken J 13:157–161. Pubmed
[14] Sasaki K, Higashihara M, Inoue K, Igarashi Y, Makino S (1976) Studies on the development of a live attenuated mumps virus vaccine. I. Attenuation of the Hoshino ‘wild’ strain of mumps virus. Kitasato Arch Exp Med 49:43–52. Pubmed
[15] Galazka AM, Robertson SE, Kraiger A (1999) Mumps and mumps vaccine: a global review. Bull World Health Organ. 77:3–14. Article Pubmed
[16] Nagai T, Okafuji T, Miyazaki C, Ito Y, Kamada M, et al. (2007) A comparative study of the incidence of aseptic meningitis in symptomatic natural mumps patients and monovalent mumps vaccine recipients in Japan. Vaccine 25:2742–2747. Article Pubmed
[17] Clarke DK, Sidhu MS, Johnson JE, Udem SA (2000) Rescue of mumps virus from cDNA. J Virol 74:4831–4838. Article Pubmed
[18] Lemon K, Rima BK, McQuaid S, Allen IV, Duprex WP (2007) The F gene of rodent brain-adapted mumps virus is a major determinant of neurovirulence. J Virol 81:8293–8302. Article Pubmed
[19] Malik T, Sauder C, Wolbert C, Zhang C, Carbone KM, et al. (2007) A single nucleotide change in the mumps virus F gene affects virus fusogenicity in vitro and virulence in vivo. J Neurovirol 13:513–521. Article Pubmed
[20] Malik TH, Wolbert C, Nerret L, Sauder C, Rubin S (2009) Single amino acid changes in the mumps virus haemagglutinin-neuraminidase and polymerase proteins are associated with neuroattenuation. J Gen Virol 90:1741–1747. Article Pubmed
[21] Reyes-Leyva J, Baños R, Borraz-Argüello M, Santos-López G, Rosas N, et al. (2007) Amino acid change 335 E to K affects the sialic-acid-binding and neuraminidase activities of Urabe AM9 mumps virus hemagglutinin-neuraminidase glycoprotein. Microbes Infect 9:234–240. Article Pubmed
[22] Santos-López G, Cruz C, Pazos N, Vallejo V, Reyes-Leyva J, et al. (2006) Two clones obtained from Urabe AM9 mumps virus vaccine differ in their replicative efficiency in neuroblastoma cells. Microbes Infect 8:332–339. Article Pubmed
[23] Sauder CJ, Zhang CX, Link MA, Duprex WP, Carbone KM, et al. (2009) Presence of lysine at aa 335 of the hemagglutinin-neuraminidase protein of mumps virus vaccine strain Urabe AM9 is not a requirement for neurovirulence. Vaccine 27:5822–5829. Article Pubmed
[24] Saito H, Takahashi Y, Harata S, Tanaka K, Sano T, et al. (1996) Isolation and characterization of mumps virus strains in a mumps outbreak with a high incidence of aseptic meningitis. Microbiol Immunol 40:271–275. Pubmed
[25] Saito H, Takahashi Y, Harata S, Tanaka K, Sato H, et al. (1998) Cloning and characterization of the genomic RNA sequence of the mumps virus strain associated with a high incidence of aseptic meningitis. Microbiol Immunol 42:133–137. Article Pubmed
[26] Ito N, Takayama-Ito M, Yamada K, Hosokawa J, Sugiyama M, et al. (2003) Improved recovery of rabies virus from cloned cDNA using a vaccinia virus-free reverse genetics system. Microbiol Immunol 47:613–617. Pubmed
[27] Rubin SA, Pletnikov M, Taffs R, Snoy PJ, Kobasa D, et al. (2000) Evaluation of a neonatal rat model for prediction of mumps virus neurovirulence in humans. J Virol 74:5382–5384. Article Pubmed
[28] Afzal MA, Marsden SA, Hull RM, Pipkin PA, Bentley ML, et al. (1999) Evaluation of the neurovirulence test of mumps vaccines. Biologicals 27:43–49. Article Pubmed
[29] Maximova O, Dragunsky E, Taffs R, Snoy R, Cogan J, et al. (1996) Monkey neurovirulence test for live mumps vaccine. Biologicals 24:233–234. Article Pubmed
[30] Saika S, Kidokoro M, Aoki A, Ohkawa T (2004) Neurovirulence of mumps virus: intraspinal inoculation test in marmosets. Biologicals 32:147–152. Article Pubmed
[31] Wolinsky JS, Stroop WG (1978) Virulence and persistence of three prototype strains of mumps virus in newborn hamsters. Arch Virol 57:355–359. Article Pubmed
[32] Rubin SA, Pletnikov M, Carbone KM (1998) Comparison of the Neurovirulence of a vaccine and a wild-type mumps virus strain in the developing rat brain. J Virol 72:8037–8042. Article Pubmed
[33] Brown EG, Dimock K, Wright KE (1996) The Urabe AM9 mumps vaccine is a mixture of viruses differing at amino acid 335 of the hemagglutinin-neuraminidase gene with one form associated with disease. J Infect Dis 174:619–622. Article Pubmed
[34] Amexis G, Fineschi N, Chumakov K (2001) Correlation of genetic variability with safety of mumps vaccine Urabe AM9 strain. Virology 287:234–241. Article Pubmed
[35] Wright KE, Dimock K, Brown EG (2000) Biological characteristics of genetic variants of Urabe AM9 mumps vaccine virus. Virus Res 67:49–57. Article Pubmed
[36] Sauder CJ, Zhang CX, Ngo L, Werner K, Lemon K, et al. (2011) Gene-specific contributions to mumps virus neurovirulence and neuroattenuation. J Virol 85:7059–7069. Article Pubmed
[37] Rubin SA, Amexis G, Pletnikov M, Li Z, Vanderzanden J, et al. (2003) Changes in mumps virus gene sequence associated with variability in neurovirulent phenotype. J Virol 77:11616–11624. Article Pubmed
[38] Chambers P, Rima BK, Duprex WP (2009) Molecular differences between two Jeryl Lynn mumps virus vaccine component strains, JL5 and JL2. J Gen Virol 90:2973–2981. Article Pubmed
[39] Tanabayashi K, Takeuchi K, Okazaki K, Hishiyama M, Yamada A (1993) Identification of an amino acid that defines the fusogenicity of mumps virus. J Virol 67:2928–2931. Article Pubmed
[40] Tanabayashi K, Takeuchi K, Hishiyama M, Yamada A (1994) Effect on fusion induction of point mutations introduced into the F protein of mumps virus. Virology 204:851–853. Article Pubmed
[41] Poch O, Blumberg, BM, Bougueleret L, Tordo N (1990) Sequence comparison of five polymerases (L proteins) of unsegmented negative-strand RNA viruses: theoretical assignment of functional domains. J Gen Virol 71:1153–1162. Article Pubmed



