Journal of Cancer Research & Therapy
An International Peer-Reviewed Open Access Journal
ISSN 2052-4994


- Download PDF
- |
- Download Citation
- |
- Email a Colleague
- |
- Share:
-
- Tweet
-

Journal of Cancer Research & Therapy
Volume 1, Issue 6, August 2013, Pages 153-162
Original researchOpen Access
miRNA array analysis determines miR-205 is overexpressed in head and neck squamous cell carcinoma and enhances cellular proliferation
-
Howard JD1,
Cheng H1,
Perez J1,
Ratner E2,
Fertig EJ1,
Considine M1,
Ochs MF1,
Slebos RJ4,
Weidhaas J2,3 and
Chung CH1,*

- 1 Department of Oncology, Sidney Kimmel Cancer Center, Johns Hopkins Medical Institute, Baltimore, MD, USA
- 2 Division of Gynecological Oncology, Yale University School of Medicine, New Haven, CT, USA
- 3 Department of Therapeutic Radiology, Yale University School of Medicine, New Haven, CT, USA
- 4 Department of Cancer Biology, Vanderbilt-Ingram Cancer Center, Vanderbilt University, Nashville, TN, USA
*Corresponding author: Chung CH, Department of Oncology, Johns Hopkins Medical Institute, 1650 Orleans Street, CRB-1 rm. 344, Baltimore, MD 21231, USA, Tel.: +410 614-6204; Fax: +410 502-0677. E-mail: cchung11@jhmi.edu
Received 23 May 2013 Revised 16 July 2013 Accepted 23 July 2013 Published 29 July 2013
DOI: http://dx.doi.org/10.14312/2052-4994.2013-24
Copyright: © 2013 Howard JD, et al. Published by NobleResearch Publishers. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
AbstractTop
MicroRNAs (miRNAs) play a critical role in cell cycle and pro-survival signal regulation. Consequently, their deregulation can enhance tumorigenesis and cancer progression. In the current investigation, we determined whether cancer- or human papillomavirus (HPV)-specific miRNA deregulation could further elucidate signal transduction events unique to head and neck squamous cell carcinoma (HNSCC). Twenty-nine newly diagnosed HNSCC tumors (HPV-positive: 14, HPV-negative: 15) and four normal mucosa samples were analyzed for global miRNA expression. Differential miRNA expression analysis concluded HNSCC is characterized by a general upregulation of miRNAs compared to normal mucosa. Additionally, miR-449a and miR-129-3p were statistically significant miRNAs differentially expressed between HPV-positive and HPV-negative HNSCC. The upregulation of miR-449a was also validated within an independent dataset obtained from TCGA containing 279 HNSCCs and 39 normal adjacent mucosa samples. To gain a better understanding of miRNA-mediated cell cycle deregulation in HNSCC, we functionally evaluated miR-205, a transcript upregulated in our cancer-specific analysis and a putative regulator of E2F1. Modulation of miR-205 with a miRNA mimic and inhibitor revealed miR-205 is capable of regulating E2F1 expression in HNSCC and overexpression of this transcript enhances proliferation. This study demonstrates miRNA expression is highly deregulated in HNSCC and functional evaluations of these miRNAs may reveal novel HPV context dependent mechanisms in this disease.
Keywords: microRNA; HPV; p16; E2F1; head and neck squamous cell carcinoma (HNSCC)
IntroductionTop
Head and neck squamous cell carcinoma (HNSCC) and its associated morbidities remain a worldwide public health concern. Traditionally, the development of HNSCC has been linked to long-term tobacco and alcohol consumption, resulting in an older patient population [1]. Recently, high-risk human papillomavirus (HPV) infection has also been described as a causal agent for HNSCC development, although this subtype affects a younger population [2]. While mutation and gene expression studies have uncovered molecular heterogeneity among HNSCCs [3-6], a decisive role for microRNAs (miRNAs) in HNSCC development is currently unclear. Thus, we wanted to determine the degree of miRNA deregulation in newly diagnosed HNSCC tumor samples compared to normal mucosa. Additionally, our tumor cohort contained equivalent numbers of HPV-positive and –negative tumors, thus we could also accurately detect miRNA deregulation specific to HPV-related disease.
miRNAs are novel global regulatory RNAs, originally discovered in C. elegans, capable of controlling hundreds of mRNA transcripts. These short, non-coding RNAs inhibit target protein expression through complimentary base-pairing with mature mRNA transcripts which blocks translation. Recent studies from several laboratories have demonstrated that miRNAs represent a critical component of human cancer development, with evidence suggesting that miRNAs regulate key proteins involved in pro-survival pathways [7]. In addition, miRNA function is extremely important for defining response to radiation therapy, chemotherapy, and can predict patient outcome in certain cancers. miRNA expression can also be altered by viral infection, as virus-specific miRNA profiles have been observed in tumors initiated by hepatitis C [8] or Epstein-Barr virus infection [9]. High-risk HPV is a sexually-transmitted virus long established as the primary factor in cervical cancer tumorigenesis. Previous work from our laboratory has shown that HPV-positivity results in a distinct global gene expression pattern apart from HPV-negative head and neck cancer patients [5]. Thus, we sought to determine whether these observations extend to miRNA expression as well, potentially explaining some of the notable proteomic differences between these two HNSCC subtypes [10].
For example, p16 is an important tumor suppressor and cell cycle regulator commonly lost in HPV-negative HNSCC [2, 11]. Yet, this protein is upregulated in HPV-positive HNSCC. In the latter case, HPV E7 initiates the targeted destruction of Rb which leads to Rb/E2F1 dysfunction and an upregulation of p16. Consequently, p16 has served as a prognostic and histologic surrogate marker for HPV infection. This subset of HNSCC is associated with the oropharynx and better overall survival [2, 12, 13]. Despite this well-studied mechanism, 4-37% of p16-negative HNSCC is HPV-positive and 0-57% of p16-positive HNSCC is HPV-negative. The discrepancies in reporting depend on the modality utilized to detect HPV and the proportion of oropharynx vs. non-oropharynx subsites within the study populations. This suggests further Rb/E2F deregulation is important in HNSCC separate from its association with HPV infection. Consequently, we were interested in determining whether candidate microRNAs could be identified within our screen that could further resolve the discrepancies observed in HNSCC cell cycle deregulation.
Previous investigations in our laboratory contributed to a better understanding of p16INK4A and Rb-associated gene function in HNSCC development [5, 10], but the involvement of miRNAs in this process is not currently understood. In the present investigation, we examined the global deregulation of miRNAs that occur in HNSCC compared to normal mucosa. In doing so, we catalog the differences in miRNA expression between HPV-positive and -negative tumors and search for potential miRNAs that may lead to a better understanding of Rb/E2F pathway deregulation in this disease.
Materials and methodsTop
Clinical specimens
Thirty-one frozen tumors from patients with newly diagnosed primary HNSCC and four normal mucosa samples (routine tonsillectomy) were collected with written informed consent under an Institutional Review Board (IRB) Health Sciences Committee (HSC)-approved protocol at Vanderbilt University Medical Center. Clinical data were obtained from Vanderbilt University Head and Neck Cancer Tissue and Clinical Database as previously described [5].
RNA isolation, RT-PCR, and DNA microarray analyses
Each tumor was examined by H&E staining to ensure presence of tumor and enriched by macro-dissection to achieve at least 70% tumor cellularity in each preparation. Total RNA was isolated using PureLink RNA Isolation Kit, and miRNA was isolated using PureLink microRNA Isolation Kit (Invitrogen, Carlsbad, CA) using approximately 10-20 milligrams of wet tissue for each sample. Quality and quantity of RNA was determined using the Agilent RNA 6000 NanoLabChip kit and Agilent 2100 bioanalyzer (Agilent Technologies, Santa Clara, CA). miRNA was utilized for the ABI Megaplex protocol without pre-amplification (Applied Biosystems, Foster City, CA). A reaction was also run using the ABI miRNA Reverse Transcription kit (#4366596) and Megaplex RT Human Pool A primers (#4399966). cDNA was then used in Taqman Universal PCR Master Mix without UNG, 2X (#4324018) per the manufacturer’s protocol. Samples were loaded onto TaqMan Low-Density Array (TLDA) cards. TLDAs were analyzed on ABI7900HT real-time PCR machine and run with the ABI default TLDA protocol. Data were normalized based on the TLDA card control.
HPV detection and DNA sequencing
Tumor DNA was isolated and HPV DNA was detected using a previously established PCR-based method [14]. Briefly, degenerate PCR primers (MY09 and MY11, WD72/76 and WD66/67/154) were used to detect highly conserved HPV L1 and E6 sequences present in all major HPV variants. Samples determined to be HPV-positive by this method were further confirmed using HPV16-specific PCR for E7 as previously described [5].
Cell culture and miRNA transfection
93VU147T (a gift from Dr. Renske D.M. Steenbergen, Vrije University, The Netherlands) and SCC15 (ATCC) HNSCC cell lines were cultured as previously described [15]. miR-205 mimic, inhibitor, and appropriate transfection controls (1µg/6-well, Dharmacon, Lafayette, CO) were transiently transfected for 6 hrs in Opti-Mem (Gibco, Grand Island, NY) using Lipofectamine 2000 (Invitrogen, Grand Island, NY). Cells were incubated in antibiotic-free medium overnight and harvested for assay or Western blot analysis the following day.
E2F1 3’UTR Luciferase assay
Cells were transfected as above with mimic and inhibitor constructs and controls. These transfections also included the E2F1 miTarget™ microRNA 3’UTR luciferase construct (1 µg/well, GeneCopoeia, Rockville, MD) and pRenilla (20 ng/well) as a transfection control. Cells proliferated for 48 hrs, were snap frozen, then lysed and analyzed for luminescence using the Dual Luciferase Reporter Assay System (Promega, Madison, WI) with the GloMax Multi Detection System (Promega, Madison, WI).
Western blot analysis
miR-205 transiently transfected cells were seeded at equivalent densities in full serum (10% FBS), allowed to proliferate for 24 hrs, and processed for SDS-PAGE and Western blot analysis as previously described [15]. Antibodies for E2F1, cleaved and total caspase-3, cyclin D1, Bim, and GAPDH were purchased from Cell Signaling Technology (Danvers, MA).
Growth assays
Transiently transfected cells were seeded in 96-well plates for Matrigel colony formation and proliferation assays in 1% serum as previously described [15]. To achieve optimal colony detection and due to the different growth rates of the cell lines, SCC15 was incubated on Matrigel (BD Biosciences, San Jose, CA) for 4 days, while 93VU147T was cultured for 6 days. Matrigel colonies were stained by MTT metabolism [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium] and quantified for colony number and average colony size using GelCount™ (Oxford Optronix, Oxford, UK) per manufacturer’s instructions. Colony diameter detection range was 20-500µm. For 96-well proliferation assays, MTS [3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium, Promega, Madison, WI] was diluted in DPBS (Dulbecco’s phosphate buffered saline, Invitrogen, Carlsbad, CA) per manufacturer’s instructions. Transfected cells were seeded in 96-well plates for 48 hrs and 20µl of MTS (1mg/mL) was added to 100µl of media in the wells. After 2-4 hrs of incubation at 37°C, 100µl of reactant was removed from the wells and absorbance was analyzed at 450nm using the GloMax Multi Detection System plate reader.
Data analysis and statistical consideration
RT-PCR miRNA data was normalized by subtracting the Ct counts of endogenous control gene expression (MammU6) from the raw Ct counts to obtain ΔCt values [16, 17]. Two samples (519CC26, 519CC74) were removed from future analyses due to overall lack of miRNA expression or MammU6 expression. In the remaining samples, transcripts determined not detectable were replaced with a maximum Ct value of 40 prior to calculating ΔCt values. Replicate ΔCt measures for one patient (519CC63 and 519CC64) were averaged for subsequent analyses. To determine differential miRNA expression, empirical Bayes moderated t-statistics were used to assess differential expression of miRNAs associated with HNSCC or HPV status. The Benjamini-Hotchberg correction [18] was applied to the resulting p-values to account for multiple hypothesis testing. All analyses were performed with the LIMMA Bioconductor package [19] implemented in R. Although samples were spread across five arrays, a batch effect was not incorporated in our statistical model because all four normal samples were analyzed on the same array. All analyses are documented in a Sweave report and included as Supplemental File 1. For miRNA family analysis, families were identified from miRBase [20-22] (miRBase.db, version 0.7.0). To cross-reference the RT-PCR data with the miRNA family database, the probes from the miRNA array were mapped to the miRBase names. Probes representing 3p or 5p miRNAs were assigned to a single name in the database. Additionally, single probes with multiple genomic loci (i.e. hsa-miR-9-1, has-miR-9-2, hsa-miR-9-3) were mapped to these appropriate locations. We also analyzed independent HNSCC miRNA expression determined with high throughput sequencing from the Cancer Genome Atlas (TCGA) for additional transcript validation [23]. HPV status was established using a combination of p16 staining and confirmation with next generation sequencing. Read per megabase mapped (RPPM) expression values were log transformed for normality [24], and differential expression was assessed using univariate, one-sided t-statistics. To avoid platform bias, analyses were performed separately for samples measured with the Illumina Genome Analyzer (7 HPV-positive and 28 HPV-negative) and Illumina HiSeq (39 normal, 28 HPV-positive, 216 HPV-negative).
ResultsTop
Patient characteristics and human papillomavirus detection
Thirty-one tumors from newly diagnosed HNSCC patients were assayed for HPV status by PCR detection of HPV viral DNA. Following RNA isolation and microarray analysis, two tumors were removed from the data set due to inadequate transcript levels or lack of MammU6 expression, the reference transcript utilized for normalization (Supplementary Figures S1A, S1B). Of the remaining tumors, 14 were HPV-positive and 15 were HPV-negative (Table 1). As expected, all 14 HPV-positive tumors were associated with the oropharynx (p=0.002, Fisher’s Exact test, oropharynx versus non-oropharynx) [25]. Additionally, the number of samples representing different clinical stages of HNSCC was equivalent between the HPV-positive and HPV-negative patient populations.
| Characteristic | HPV (+) N=14 | HPV (-) N=15 | Total N=29 | p-values |
| Age (median, range) | 53.5 (38-73) | 58 (30-77) | 55.5 (30-77) | 0.651 |
| Sex | ||||
| Male | 14 | 13 | 27 | |
| Female | 0 | 2 | 2 | 0.482 |
| Race | ||||
| White | 14 | 12 | 26 | |
| Black | 0 | 2 | 2 | |
| Other | 0 | 1 | 1 | 0.352 |
| Tumor sites | ||||
| Oral cavity | 0 | 5 | 5 | |
| Oropharynx | 14 | 7 | 21 | |
| Larynx | 0 | 2 | 2 | |
| Hypopharynx | 0 | 1 | 1 | 0.0023 |
| Clinical stage | ||||
| I-II | 1 | 2 | 3 | |
| III | 3 | 3 | 6 | |
| IV | 10 | 10 | 20 | 0.664 |
| Tumor Grade | ||||
| Well differentiated | 0 | 3 | 3 | |
| Moderately differentiated | 7 | 7 | 14 | |
| Poorly differentiated | 7 | 5 | 12 | 0.134 |
| Tobacco use | ||||
| Never | 5 | 1 | 6 | |
| Ever | 9 | 14 | 23 | 0.082 |
1Wilcoxon Mann-Whitney Rank test
2Fisher’s Exact test
3Fisher’s Exact test, oropharynx vs. others
4X2 test for trend.
miRNA expression is globally upregulated in HNSCC compared to normal mucosa
Following normalization, tumor-associated miRNA expression was analyzed for differential transcripts compared to normal mucosa. The ΔCt values for tumor-associated miRNAs demonstrated a much broader expression profile than normal tissue (Supplementary Figure S2), suggesting a less restrictive miRNA profile is associated with HNSCC. When the differential expression of specific miRNA transcripts was analyzed, 154 miRNAs were significantly upregulated while 103 were concomitantly downregulated in the tumor versus normal analysis (Benjamini-Hotchberg adj. p-values <0.05) (Supplementary Tables S1, S2). The total number and degree to which the individual transcripts are significantly modulated indicates miRNAs are generally upregulated in HNSCC compared to normal tissue. One striking exception to this observation is the robust downregulation of miR-124 (ΔΔCt: -9.27, adj. p-value: 2.0x10-7). This miRNA is an important tumor suppressor which is silenced by hypermethylation in a number of different cancers [26-28]. Consequently, this gene may experience similar epigenetic silencing in HNSCC.
To gain further insight from the differential transcript analysis, deregulated miRNAs from our screen were grouped according to miRBase family designations (miRBase.org). When the differential transcripts were organized in this fashion, seven families of miRNAs were significantly upregulated while two were downregulated (Table 2, Supplementary Table 3). Once again, the magnitude and sum total of miRNA deregulation favored an upregulation of critical miRNA families. Upregulated families of note include genes classically associated with the development of cancer: let-7 (MIPF0000002), miR-200 (MIPF0000019), and members of the miR-17-92 and miR-106b-25 clusters (MIPF0000001). While these upregulated families have been studied intensively, the function of the downregulated families remains poorly understood and may represent novel tumor suppressors regulating HNSCC-specific oncogenes.
| miRBase family ID | Family members on microarray | Members detected | microRNAs |
| Upregulated | |||
| MIPF0000002 | 8 | 8 | let-7b, let-7g, let-7d, let7f, let-7a, let-7e, miR-98, let-7c |
| MIPF0000001 | 8 | 7 | miR-20a, miR-18a, miR-17, miR-106a, miR—93, miR-106b, miR-20b, miR-18b |
| MIPF0000019 | 5 | 5 | miR-200a, miR-429, miR-141, miR-200c, miR-200b |
| MIPF0000006 | 4 | 4 | miR-15b, miR-16, miR-195, miR-15a |
| MIPF0000034 | 4 | 4 | miR-130b, miR-301a, miR-130a, miR-301b |
| MIPF0000009 | 3 | 3 | miR-29b, miR-29a, miR-29c |
| MIPF0000056 | 3 | 3 | miR-148a, miR-148b, miR-152 |
| Downregulated | |||
| MIPF0000020 | 33 | 18 | miR-519c-3p, miR-523, miR-518d-3p, miR-518d-5p, miR-525-5p, miR-525-3p, miR-520b, miR-518a-5p, miR-520d-5p, miR-524-5p, miR-520e, miR-521, miR-516a-5p, miR-520a-5p, miR-520a-3p, miR-526b, miR-520f, miR-519e |
| MIPF0000130 | 11 | 8 | miR-512-5p, miR-512-3p, miR-507, miR-506, miR-509-5p, miR-509-3-5p, miR-510, miR-508-3p, miR-508-5p, miR-513-5p, miR-511 |
To determine whether specific miRNA profiles were associated with HPV status, HPV-positive miRNA expression was compared to the HPV-negative transcript levels. Only two miRNAs achieved statistical significance for differential expression between these two groups: miR-449a was upregulated (ΔΔCt: 2.76, adj. p-value: 0.0262) and miR-129-3p was downregulated (ΔΔCt: -3.82, adj. p-value: 0.0262) in HPV-positive versus HPV-negative HNSCC (Supplemental Figure S3). The average expression of both miRNAs was more similar to normal tissue in HPV-negative samples suggesting the mechanisms for the differential expression are specific to HPV infection.
To further explore these findings in other dataset, HPV-specific differential expression of these miRNAs was evaluated within the HNSCC samples available in The Cancer Genome Atlas (TCGA) [23]. In the larger data set (28 HPV-positive vs. 216 HPV-negative, Illumina HiSeq), miR-449a was confirmed as a transcript upregulated in HPV-positive HNSCCs (p=0.02, Supplemental Figure S4). miR-449a trended towards significance in the tumors analyzed with Illumina GA (p=0.08); however, this was in a much smaller dataset (7 HPV-positive vs. 28 HPV-negative). Unfortunately, the miRNA sequencing reads in this dataset were not aligned to miR-129-3p in the level 3 data. In light of recent genomic sequencing projects and the mounting clinical evidence that these subtypes of HNSCC represent very different diseases [2-4], the degree of similarity they share on a miRNA level is unexpected.
miR-205 can regulate E2F1 expression in HNSCC cell lines
Of the miRNAs differentially expressed between HNSCC and normal mucosa in our screen, we were interested in characterizing HNSCC-specific miRNAs that may further deregulate cell cycle protein expression and function. After reviewing the putative genes within our dataset, it was noted that miR-205 (ΔΔCt: 2.76, adj. p-value: 0.0262) is a putative regulator of the Rb/E2F pathway by directly modulating E2F1 protein expression [30]. To further validate miR-205 is upregulated in HNSCC, this transcript was analyzed for differential expression within the Illumina HiSeq dataset containing 28 HPV-positive and 216 HPV-negative HNSCCs as well as 39 adjacent normal samples (Supplemental Figure S5). Once again, miR-205 was overexpressed in HPV-positive and –negative HNSCC with respect to normal control (p-value=1.2 x 10-5).
miRNA regulation of protein targets is highly context dependent and characterization needs to occur within the specific genetic context of interest. Consequently, to validate miR-205 regulation of E2F1 in HNSCC cell lines, an HPV-positive (93VU147T) and HPV-negative (SCC15) cell line was transiently transfected with a miR-205 mimic, inhibitor, or appropriate transfection control. These cells were also co-transfected with a luciferase reporter regulated by the 3’UTR of E2F1 (Figure 1A). miR-205 mimic and inhibitor demonstrated appropriate suppression and upregulation of luciferase protein in both cell lines confirming E2F1 is a direct target of miR-205 regulation in HNSCC.
To further establish that miR-205 can regulate E2F1 protein expression in HNSCC, cells were transfected as before and modulations in E2F1 and additional members of Rb/E2F signaling and apoptotic signaling pathways were assayed by Western blot (Figure 1B). In the HPV-positive cell line (93VU147T), miR-205 status demonstrated a substantial inverse correlation with E2F1 protein expression (Figure 1B). miR-205 suppression of E2F1 was also associated with an upregulation of cyclin D1 and decreased expression of pro-apoptotic signaling members (cleaved caspase-3, Bim), suggesting E2F1 may have a tumor suppressor role in 93VU147T.
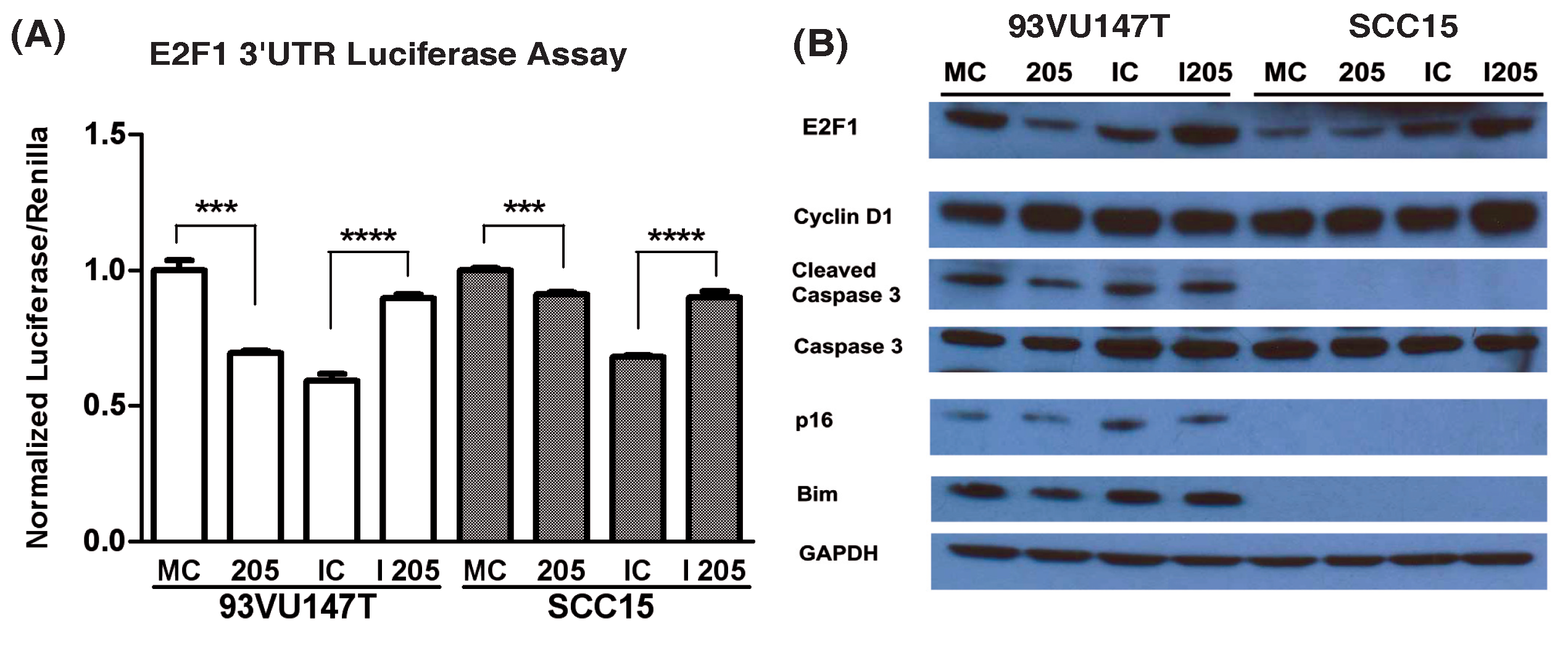
Figure 1 miR-205 modulates E2F1 expression in HNSCC cell lines. 93VU147T and SCC15 were co-transfected with mimic control (MC), miR-205 (205), inhibitor control (IC), or miR-205 inhibitor (I 205) and a luciferase construct regulated by the E2F1 3’UTR. Cells were passively lysed and analyzed (A) for E2F1 3’UTR activity by chemiluminescent detection of luciferase and renilla transfection control. 93VU147T and SCC15 were then transiently transfected with the same miRNA modulatory constructs and allowed to proliferate 24 hrs before SDS-PAGE and Western blot analysis (B) for changes in target and downstream protein expression. ***p<0.001, ****p<0.0001.
However, in SCC15 miR-205 regulation of E2F1 appears slightly different. miR-205 overexpression did not have a significant effect on E2F1 expression while miR-205 inhibition increased E2F1 levels. Additionally, cleaved caspase-3, Bim, and p16 were not detected in this cell line, common consequences of HPV-negative HNSCC development. Consequently, the loss of these proteins could affect the regulation and functional role of E2F1 in this context.
miR-205 overexpression increases HNSCC cell line growth
To determine the functional effects of miR-205 modulation in HNSCC, 93VU147T and SCC15 were transiently transfected with the same miR-205 constructs as before and changes in growth were assayed. Cells were seeded in 96-well format and changes in proliferation were estimated by MTS metabolism (Figure 2A). miR-205 overexpression increased proliferation while miR-205 inhibition decreased proliferation in these cells, with the most significant growth inhibition occurring in SCC15. To determine how miR-205 modulation affects three-dimensional cell growth, transiently transfected cells were seeded on Matrigel and allowed to proliferate 4-6 days in an anchorage-independent manner. SCC15 recapitulated the more aggressive phenotype reflective of an HPV-negative cell line lacking p16 and pro-apoptotic pathway members (Figure 2B). This cell line was more clonigenic and produced larger colonies than the HPV-positive counterpart (93VU147T). Once quantified, we determined miR-205 overexpression induced a marked increase average colony area, while SCC15 again experienced the most significant decrease in growth with miR-205 inhibition (Figure 2C). Thus, the three-dimensional growth characteristics recapitulated the two-dimensional growth modulation previously observed.
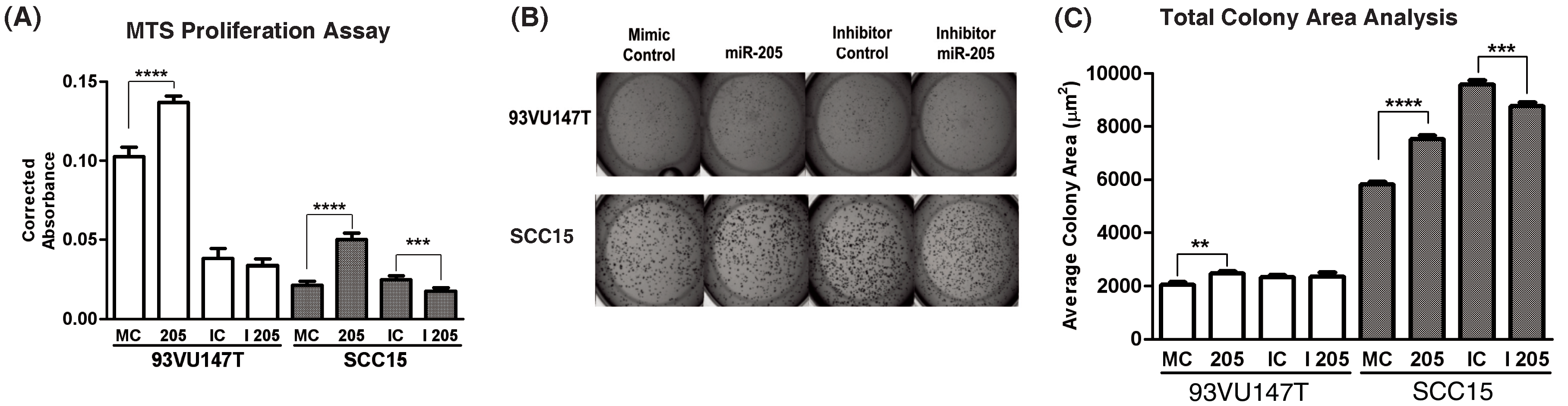
Figure 2 miR-205 overexpression enhances HNSCC cell line growth in vitro. 93VU147T and SCC15 were transiently transfected with mimic control (MC), miR-205 (205), inhibitor control (IC), or miR-205 inhibitor (I 205) as before and growth was estimated by MTS metabolism (A) after 2 days of growth in 1% serum. These cells were also seeded in Matrigel colony formation assays (B) to determine changes in clonigenicity and proliferation during anchorage-independent growth. Quantification of average colony size (C, mean with SEM) per condition. **p<0.01, ***p<0.001, ****p<0.0001.
Consequently, miR-205 is a potential HNSCC oncogene capable of enhancing proliferation in an HPV-independent manner. This observation is consistent with the fact this transcript was upregulated in both HPV-positive and HPV-negative tumors in our miRNA analysis. Thus, miR-205 upregulation can alter proliferation in both cell lines, but the specific phenotypic modulations may depend on the degree of E2F1 modulation and its function within its associated molecular landscape.
DiscussionTop
To assess miRNA deregulation in HNSCC, we catalogued the differentially expressed miRNAs between malignant tumors and normal mucosa. Additionally, we also determined the HPV-specific differential miRNA expression within our tumor cohort. The subsequent analysis has determined that HNSCCs experience a net gain of transcripts compared to normal mucosa, with a few interesting exceptions. Similar miRNA investigations in HNSCC have also observed a net gain in miRNAs rather than a loss [31-34]. Although the degree to which individual transcripts are upregulated differs widely between these studies, the general trends in miRNA deregulation are concordant. These results imply two possibilities: (1) miRNAs classically studied as tumor suppressors mediate oncogenic events in HNSCC, or (2) HNSCC development is uniquely reliant on tumor suppressor loss and a net gain of miRNAs could exacerbate this process.
In the first instance, a closer investigation of the families significantly upregulated within our analysis could yield some insight. For example, the let-7 family (MIPF0000002) is widely considered to exert tumor suppressive functions through its negative regulation of Ras and HMGA2 [35-37]. However, let-7b is specifically downregulated in acute lymphoblastic leukemia [38], melanoma [39], and ovarian cancer [40], but upregulated in gastric cancer [41] and our current analysis. Let-7f is also downregulated in sarcoma [42], lung [43], ovarian [44], and prostate cancer [45], while it is upregulated in breast cancer [46] and our current study. Furthermore, let-7i was upregulated in a previous HNSCC screen [31], however this transcript was not statistically significant within our analysis. Consequently, the end result of miRNA deregulation is very likely context dependent. The equilibrium of hundreds of target transcripts unique to different tissue types defines the pathological consequences of miRNA modulation. Thus, it is imperative that the function of these families is rigorously evaluated in future HNSCC investigations to determine their decisive role in contributing to this specific disease.
In the second case, recent sequencing projects have suggested tumor suppressor loss is a major contributing factor to the development of HNSCC, despite the heterogeneity of the disease [3, 4]. These mutations are more common in HPV-negative disease, simply because HPV viral proteins actively disable the p53 and pRb pathways to similar effect, rendering the requirement for additional tumor suppressor mutation redundant. Previous studies have also established that HNSCC is associated with a hypomethylated genome compared to normal mucosa [47]. Thus, it could be expected that hypomethylation may drive the net gain of miRNA expression observed in HNSCC. In other cancers, the hypothesis that miRNA downregulation leads to tumorigenesis hinges on the likelihood that repression of candidate oncogenes is relieved during deregulation. However, active targeting of tumor suppressor pathways by miRNA overexpression may also represent a viable means of deregulating growth and cell cycle checkpoints. Although miRNAs can regulate innumerable targets, the probability of tumor suppressor loss increases as the number of upregulated miRNAs accumulate.
The sheer number of miRNAs deregulated in our tumor versus normal analysis is substantial and the power of this analysis is admittedly hampered by a statistical limitation. Differential expression analysis between a large number of tumors (n=29) and a small number of normal samples (n=4) increases the probably a given transcript will experience a statistically significant modulation. Thus, it is possible the sum total of significant transcripts is artificially inflated due to a lack of statistical power or batch effects. Nonetheless, the observed trend in miRNA upregulation is also recapitulated within the miRBase family analysis, reliant on groups of miRNAs acting in concert rather than individual genes. As mentioned above, while miRNA family modulation can vary between tumor subtypes, the let-7 (MIPF0000002), miR-17-92 (MIPF0000001), and miR-200 (MIPF0000019) families have been intensely studied and represent critical modifiers of cancer-associated signal transduction.
Although the aforementioned statistical considerations hampered the power of tumor versus normal analyses, this was not a concern when comparing miRNA expression between HPV-positive (n=14) and HPV-negative (n=15) tumors which were randomized across three separate microarray plates. Only two miRNAs were differentially expressed between these groups (miR-449a ΔΔCt: 2.76, miR-129-3p ΔΔCt: -3.82; HPV-positive vs. HPV-negative). This was a surprising result given the myriad ways HPV-positive and -negative tumors differ in gene and protein expression [5, 10], genomic mutational profiles [3, 4], and clinical outcome [2]. Based on previous investigations, there are known mechanistic consequences of miR-449a and miR-129-3p deregulation. Notch1 is a validated target of miR-449a and a putative tumor suppressor in HNSCC [3, 4, 48]. Additionally, Notch2 and two Notch ligands (DLL1 and JAG1) have been computationally defined as targets of miR-449a [30]. Thus, overexpression of this transcript could serve to suppress Notch signaling by simultaneously downregulating the receptors and ligands associated with the activity of this critical pathway. This would also provide a selective advantage for HPV-positive HNSCC development as Notch1 downregulation is required for sustained E6/E7 expression and transformation [49].
While the function of miR-129-3p itself is largely unknown, its precursor transcript, miR-129-2, is a well-studied tumor suppressor and frequently hypermethylated in a number of different cancers [29, 50-52]. A major consequence of miR-129-2 silencing is SOX4 upregulation [29], a feature that has not yet been described in the HPV-positive HNSCC proteome. However, recent evidence suggests that the transfection of immortalized keratinocytes (HaCaT) with various HPV genomes is capable of specifically downregulating miR-129-3p, the fully-mature processed transcript observed in our study [53]. Future investigations will be required to determine whether HPV-specific miR-129-3p silencing involves epigenetic mechanisms or more traditional means of gene expression regulation. More importantly, differential miRNA expression is not a likely explanation for the vast differences observed between these subtypes of disease. Consequently, we sought to investigate shared putative genes between these subtypes that may yield more clinically relevant mechanisms for general HNSCC deregulation.
Prior work in our laboratory elucidated the deregulation of cell cycle pathways in HNSCC and we wanted to extend these findings within our differential miRNA transcripts. Consequently, we evaluated miR-205 as a putative regulator of E2F1 expression and associated pathway signaling. Prior investigations of miRNAs in HNSCC uncovered a co-upregulation of the miR-200 family and miR-205 in CD44low cells [54], both of which are overexpressed in our current analysis. However, the explicit function of miR-205 was not assayed in this study. This transcript has also been utilized as an exquisitely sensitive diagnostic marker for HNSCC detection in lymph node metastases [55], however our current analysis represents the first time this gene had been functionally evaluated in the context of this specific disease. In a manner analogous to HNSCC let-7 upregulation, this gene set represents a miRNA family classically studied as a tumor suppressor which is paradoxically upregulated in this disease [56]. While putative targets of the miR-200 family have received much more attention, little is known concerning miR-205-mediated HNSCC signaling. Previous studies have demonstrated that miR-205 can be an oncogene or a tumor suppressor depending on the genetic context. For example, miR-205 is overexpressed in cervical cancer and functionally serves to increase the proliferation and migration of these cells [57]. However, miR-205 is generally observed to have tumor suppressive functions. miR-205 is specifically down regulated in prostate cancer and is associated with epithelial-to-mesenchymal transition and docetaxel resistance [58, 59]. Additionally, high expression of E2F1 in malignant melanoma has been shown to downregulate miR-205 and lead to anticancer drug resistance [60]. Consequently, the function of miR-205 needs to be evaluated within each specific genetic context.
Using two-dimensional and three-dimensional growth assays, we have demonstrated that miR-205 overexpression increases the proliferation of HNSCC cell lines independent of HPV status. After 3’UTR and target protein analysis, we determined miR-205 is capable of regulating E2F1 expression in HPV-positive and HPV-negative HNSCC cells. However, the modulations that occur downstream may be functionally dependent on HPV status despite sharing a similar phenotypic outcome. A recent proteomic screen of HPV-positive and HPV-negative tumors determined an E2F1 gene signature is associated with the protein expression profile of HPV-positive tumors [10]. Consequently, HPV-positive cells may have a unique sensitivity to modulations in E2F1 protein expression not inherent to HPV-negative cells. Indeed, 93VU147T demonstrated the most significant E2F1 modulations following miR-205 manipulation. However, similar functional changes occurred in SCC15 following miR-205 modulation without consistent changes in E2F1 protein. Aside from a lack of apoptotic pathway and p16 expression, SCC15 is also characterized by high levels of EGFR and AKT activation [61]. Previous studies have demonstrated that activated PI3K/AKT signaling is capable of repressing genes which cooperate with E2F1-mediated apoptosis [62] . Thus, miR-205 modulation of E2F1 in SCC15 may be tangential to its oncogenic effects due to additional targets unique to its molecular context. It was already well-established that miRNAs can have differential functions depending on the disease-state in question, and our current investigation has determined that the same miRNA can have differential signaling consequences within the same disease. However, the paths utilized depend on the subtype and genetic context available, yet ultimately these changes result in similar phenotypic modulations. Consequently, our miRNA screen and functional studies suggest miR-205 can enhance the HNSCC oncogenic phenotype regardless by multiple molecular mechanisms independent of HPV status.
ConclusionTop
We have determined that miRNAs are globally upregulated in human HNSCCs and only modest differences exist in the miRNA profile defining HPV-positive and HPV-negative disease. To begin characterizing miRNAs of interest within our screen, we functionally evaluated miR-205 in an HPV-positive and HPV-negative cell line. We determined miR-205 is capable of regulating E2F1 protein expression in HNSCC and this transcript classically studied as a tumor suppressor provides a growth advantage to both cell lines. Yet, the specific mechanisms inherent to miR-205 HNSCC oncogenic signaling may depend on the molecular context. Consequently, our functional evaluation has determined that cancer-specific miRNA deregulation can have similar phenotypic outcomes despite relying on HPV context dependent mechanisms. Thus, future investigations of the miRNAs reported within this analysis will serve as excellent examples of context dependent signaling in HNSCC and potentially reveal critical signaling interactions targetable by pharmacological intervention.
Acknowledgements
The project was funded in part by Damon Runyon Clinical Investigator Award (CL-28-05) and NIH NIDCR R01 (DE017982) to CHC. This work was also supported in part by the Radiation Therapy Oncology Group Translational Research Program, funded through RTOG U10 CA21661 and CCOP U10 CA37422 from the National Cancer Institute (NCI). The contents of this article are the sole responsibility of the authors and do not necessarily represent the official views of the NCI or the Commonwealth of Pennsylvania.
Disclosure
The authors declare they have no competing interests as defined by Journal of Cancer Research and Therapy, or other interests that might be perceived to influence the results and discussion reported in this paper.
Supplementary data
Analysis of miRNA data in HNSCC (Elana J. Fertig)
Supplementary figures
Supplementary tables
ReferencesTop
[1] Forastiere A, Koch W, Trotti A, Sidransky D (2001) Head and neck cancer. N Engl J Med 345:1890–1900. Article Pubmed
[2] Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, et al. (2010) Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med 363:24–35. Article Pubmed
[3] Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, et al. (2011) Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science 333:1154–1157. Article Pubmed
[4] Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, et al. (2011) The mutational landscape of head and neck squamous cell carcinoma. Science 333:1157–1160. Article Pubmed
[5] Slebos RJ, Yi Y, Ely K, Carter J, Evjen A, et al. (2006) Gene expression differences associated with human papillomavirus status in head and neck squamous cell carcinoma. Clin Cancer Res 12:701–709. Article Pubmed
[6] Chung CH, Parker JS, Karaca G, Wu J, Funkhouser WK, et al. (2004) Molecular classification of head and neck squamous cell carcinomas using patterns of gene expression. Cancer Cell 5:489–500. Article Pubmed
[7] Lujambio A, Lowe SW (2012) The microcosmos of cancer. Nature 482:347–355. Article Pubmed
[8] Diaz G, Melis M, Tice A, Kleiner DE, Mishra L, et al. (2013) Identification of microRNAs specifically expressed in hepatitis C virus-associated hepatocellular carcinoma. Int J Cancer 133:816–824. Article Pubmed
[9] Motsch N, Alles J, Imig J, Zhu J, Barth S, et al. (2012) MicroRNA profiling of Epstein-Barr virus-associated NK/T-cell lymphomas by deep sequencing. PLoS One 7:e42193. Article Pubmed
[10] Slebos RJ, Jehmlich N, Brown B, Yin Z, Chung CH, et al. (2013) Proteomic analysis of oropharyngeal carcinomas reveals novel HPV-associated biological pathways. Int J Cancer 132:568–579. Article Pubmed
[11] Demokan S, Chuang A, Suoğlu Y, Ulusan M, Yalnız Z, et al. (2012) Promoter methylation and loss of p16(INK4a) gene expression in head and neck cancer. Head Neck 34:1470–1475. Article Pubmed
[12] Weinberger PM, Yu Z, Haffty BG, Kowalski D, Harigopal M, et al. (2006) Molecular classification identifies a subset of human papillomavirus--associated oropharyngeal cancers with favorable prognosis. J Clin Oncol 24:736–747. Article Pubmed
[13] Rischin D, Young RJ, Fisher R, Fox SB, Le QT, et al. (2010) Prognostic significance of p16INK4A and human papillomavirus in patients with oropharyngeal cancer treated on TROG 02.02 phase III trial. J Clin Oncol 28:4142–4148. Article Pubmed
[14] Resnick RM, Cornelissen MT, Wright DK, Eichinger GH, Fox HS, et al. (1990) Detection and typing of human papillomavirus in archival cervical cancer specimens by DNA amplification with consensus primers. J Natl Cancer Inst 82:1477–1484. Article Pubmed
[15] Hatakeyama H, Cheng H, Wirth P, Counsell A, Marcrom SR, et al. (2010) Regulation of heparin-binding EGF-like growth factor by miR-212 and acquired cetuximab-resistance in head and neck squamous cell carcinoma. PLoS One 5:e12702. Article Pubmed
[16] Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. Article Pubmed
[17] Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3:1101–1108. Article Pubmed
[18] Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society Series B 57:289–300. Article
[19] Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3: 1544–6115. Article Pubmed
[20]Griffiths-Jones S (2004) The microRNA Registry. Nucleic Acids Res 32: 109–111. Article Pubmed
[21] Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34: 140–144. Article Pubmed
[22] Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39: 152–157. Article Pubmed
[23] TCGA Research Network (2013) HNSCC Marker Paper. In submission.
[24] Law CW, Chen Y, Shi W, Smyth GK (2013) Voom! Precision weights unlock linear model analysis tools for RNA-seq read counts. In submission. Article
[25] Gillison ML (2004) Human papillomavirus-associated head and neck cancer is a distinct epidemiologic, clinical, and molecular entity. Semin Oncol 31:744–754. Article Pubmed
[26] Wong KY, So CC, Loong F, Chung LP, Lam WW, et al. (2011) Epigenetic inactivation of the miR-124–121 in haematological malignancies. PLoS One 6:e19027. Article Pubmed
[27] Furuta M, Kozaki KI, Tanaka S, Arii S, Imoto I, et al. (2010) miR-124 and miR-203 are epigenetically silenced tumor-suppressive microRNAs in hepatocellular carcinoma. Carcinogenesis 31:766–776. Article Pubmed
[28] Wilting SM, van Boerdonk RA, Henken FE, Meijer CJ, Diosdado B, et al. (2010) Methylation-mediated silencing and tumour suppressive function of hsa-miR-124 in cervical cancer. Mol Cancer 9:167. Article Pubmed
[29] Huang YW, Liu JC, Deatherage DE, Luo J, Mutch DG, et al. (2009) Epigenetic repression of microRNA-129–122 leads to overexpression of SOX4 oncogene in endometrial cancer. Cancer Res 69:9038–9046. Article Pubmed
[30] TargetScan 6.1. 2012 March 2012 Article
[31] Chang SS, Jiang WW, Smith I, Poeta LM, Begum S, et al. (2008) MicroRNA alterations in head and neck squamous cell carcinoma. Int J Cancer 123:2791–2797. Article Pubmed
[32] Lajer CB, Garnæs E, Friis-Hansen L, Norrild B, Therkildsen MH, et al. (2012) The role of miRNAs in human papilloma virus (HPV)-associated cancers: bridging between HPV-related head and neck cancer and cervical cancer. Br J Cancer 106:1526–1534. Article Pubmed
[33] Nurul-Syakima AM, Yoke-Kqueen C, Sabariah AR, Shiran MS, Singh A, et al. (2011) Differential microRNA expression and identification of putative miRNA targets and pathways in head and neck cancers. Int J Mol Med 28:327–336. Article Pubmed
[34] Lajer CB, Nielsen FC, Friis-Hansen L, Norrild B, Borup R, et al. (2011) Different miRNA signatures of oral and pharyngeal squamous cell carcinomas: a prospective translational study. Br J Cancer 104:830–840. Article Pubmed
[35] Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, et al. (2005) RAS is regulated by the let-7 microRNA family. Cell 120:635–647. Article Pubmed
[36] Park SM, Shell S, Radjabi AR, Schickel R, Feig C, et al. (2007) Let-7 prevents early cancer progression by suppressing expression of the embryonic gene HMGA2. Cell Cycle 6:2585–2590. Article Pubmed
[37] Mayr C, Hemann MT, Bartel DP (2007) Disrupting the pairing between let-7 and Hmga2 enhances oncogenic transformation. Science 315:1576–1579. Article Pubmed
[38] Mi S, Lu J, Sun M, Li Z, Zhang H, et al. (2007) MicroRNA expression signatures accurately discriminate acute lymphoblastic leukemia from acute myeloid leukemia. Proc Natl Acad Sci U S A 104:19971–19976. Article Pubmed
[39] Schultz J, Lorenz P, Gross G, Ibrahim S, Kunz M (2008) MicroRNA let-7b targets important cell cycle molecules in malignant melanoma cells and interferes with anchorage-independent growth. Cell Res 18:549–557. Article Pubmed
[40] Nam EJ, Yoon H, Kim SW, Kim H, Kim YT, et al. (2008) MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 14:2690–2695. Article Pubmed
[41] Motoyama K, Inoue H, Nakamura Y, Uetake H, Sugihara K, et al. (2008) Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clin Cancer Res 14:2334–2340. Article Pubmed
[42] Subramanian S, Lui WO, Lee CH, Espinosa I, Nielsen TO, et al. (2008) MicroRNA expression signature of human sarcomas. Oncogene 27:2015–2026. Article Pubmed
[43] Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, et al. (2004) Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64:3753–3756. Article Pubmed
[44] Dahiya N, Sherman-Baust CA, Wang TL, Davidson B, Shih IeM, et al. (2008) MicroRNA expression and identification of putative miRNA targets in ovarian cancer. PLoS One 3:e2436. Article Pubmed
[45]Ozen M, Creighton CJ, Ozdemir M, Ittmann M (2008) Widespread deregulation of microRNA expression in human prostate cancer. Oncogene 27:1788–1793. Article Pubmed
[46] Yan LX, Huang XF, Shao Q, Huang MY, Deng L, et al. (2008) MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA 14:2348–2360. Article Pubmed
[47] Smith IM, Mydlarz WK, Mithani SK, Califano JA (2007) DNA global hypomethylation in squamous cell head and neck cancer associated with smoking, alcohol consumption and stage. Int J Cancer 121:1724–1728. Article Pubmed
[48] Capuano M, Iaffaldano L, Tinto N, Montanaro D, Capobianco V, et al. (2011) MicroRNA-449a overexpression, reduced NOTCH1 signals and scarce goblet cells characterize the small intestine of celiac patients. PLoS One 6:e29094. Article Pubmed
[49] Talora C, Sgroi DC, Crum CP, Dotto GP (2002) Specific down-modulation of Notch1 signaling in cervical cancer cells is required for sustained HPV-E6/E7 expression and late steps of malignant transformation. Genes Dev 16:2252–2263. Article Pubmed
[50] Chen X, Zhang L, Zhang T, Hao M, Zhang X, et al. (2013) Methylation-mediated repression of microRNA 129–122 enhances oncogenic SOX4 expression in HCC. Liver Int 33:476–486. Article Pubmed
[51] Lu CY, Lin KY, Tien MT, Wu CT, Uen YH, et al. (2013) Frequent DNA methylation of MiR-129–122 and its potential clinical implication in hepatocellular carcinoma. Genes Chromosomes Cancer 52:636–643. Article Pubmed
[52] Tsai KW, Wu CW, Hu LY, Li SC, Liao YL, et al. (2011) Epigenetic regulation of miR-34b and miR-129 expression in gastric cancer. Int J Cancer 129:2600–2610. Article Pubmed
[53] Dreher A, Rossing M, Kaczkowski B, Andersen DK, Larsen TJ, et al. (2011) Differential expression of cellular microRNAs in HPV 11, -16, and -45 transfected cells. Biochem Biophys Res Commun 412:20–25. Article Pubmed
[54] Wiklund ED, Gao S, Hulf T, Sibbritt T, Nair S, et al. (2011) MicroRNA alterations and associated aberrant DNA methylation patterns across multiple sample types in oral squamous cell carcinoma. PLoS One 6:e27840. Article Pubmed
[55] Fletcher AM, Heaford AC, Trask DK. (2008) Detection of metastatic head and neck squamous cell carcinoma using the relative expression of tissue-specific mir-205. Transl Oncol 1:202–208. Article Pubmed
[56] Schliekelman MJ, Gibbons DL, Faca VM, Creighton CJ, Rizvi ZH, et al. (2011) Targets of the tumor suppressor miR-200 in regulation of the epithelial-mesenchymal transition in cancer. Cancer Res 71:7670–7682. Article Pubmed
[57] Xie H, Zhao Y, Caramuta S, Larsson C, Lui WO. (2012) miR-205 expression promotes cell proliferation and migration of human cervical cancer cells. PLoS One 7:e46990. Article Pubmed
[58] Puhr M, Hoefer J, Schäfer G, Erb HH, Oh SJ, et al. (2012) Epithelial-to-mesenchymal transition leads to docetaxel resistance in prostate cancer and is mediated by reduced expression of miR-200c and miR-205. Am J Pathol 181:2188–2201. Article Pubmed
[59] Verdoodt B, Neid M, Vogt M, Kuhn V, Liffers ST, et al. (2013) MicroRNA-205, a novel regulator of the anti-apoptotic protein Bcl2, is downregulated in prostate cancer. Int J Oncol 43:307–314. Article Pubmed
[60] Alla V, Kowtharapu BS, Engelmann D, Emmrich S, Schmitz U, et al. (2012) E2F1 confers anticancer drug resistance by targeting ABC transporter family members and Bcl-2 via the p73/DNp73-miR-205 circuitry. Cell Cycle 11:3067–3078. Article Pubmed
[61] Hatakeyama H, Cheng H, Fertig E, Considine M, Dahlman K, et al. (2011) Epithelial-to-mesenchymal transition and cetuximab resistance in head and neck squamous cell carcinoma. AACR 2011. Orlando, FL.
[62] Hallstrom TC, Mori S, Nevins JR (2008) An E2F1-dependent gene expression program that determines the balance between proliferation and cell death. Cancer Cell 13:11–22. Article Pubmed


